Introduction
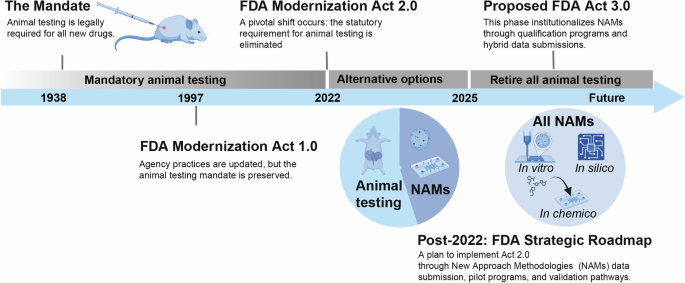
A shift is currently transforming the preclinical drug development landscape. This evolution is driven by a convergence of mounting scientific evidence and decisive legislative action, both of which are steering the industry away from its long-standing dependence on animal testing. The historical mandate for animal models, dating back to 1938, has recently faced intense scrutiny due to its poor predictive validity1. This inefficiency is underscored by an high attrition rate: over 90% of drug candidates that appear safe and effective in animal studies fail to gain FDA approval, largely due to unpredicted toxicities or a lack of efficacy in humans2,3,4,5. In response to these challenges, the scientific community has introduced New Approach Methodologies (NAMs), an human-relevant testing platforms engineered to enhance clinical translatability6,7. This movement gained definitive legislative support with the passage of the FDA Modernization Act 2.0 in 20228,9. This landmark legislation eliminated the statutory requirement for animal testing and explicitly authorized the use of NAMs to support Investigational New Drug (IND) applications. This represents a significant departure from the FDA Modernization Act of 1997, which updated agency practices but preserved the original animal testing mandates (Fig. 1).
To support this legislative evolution, a recent FDA roadmap has articulated a detailed strategy for the systematic integration of validated NAMs10. The implementation initially focuses on monoclonal antibodies (mAbs), an area where animal models have proven particularly unreliable for predicting human immunogenicity10. The strategy outlines a multi-pronged approach, including the parallel submission of NAM data alongside traditional animal studies to build a robust comparative repository10. This effort is coordinated through collaborative bodies such as the Interagency Coordinating Committee on the Validation of Alternative Methods (ICCVAM) and partners like the National Institutes of Health (NIH), with the ultimate goal of reducing animal use and fostering a regulatory environment where validated, human-centric NAMs become the standard10.
For physiological systems, such as dynamic culture, organoids, and organ-on-a-chip platforms have demonstrated distinct limitations when tasked with replicating complex systemic functions11. Many of these devices are constrained by material deficiencies, such as the absorption of hydrophobic compounds by polydimethylsiloxane (PDMS), which can distort pharmacokinetic assessments12. Furthermore, they often exhibit architectural simplicity, failing to capture the full complexity of native tissue, such as cylindrical geometries, pulsatile flow, or the presence of essential stromal cells like fibroblasts and pericytes13,14,15,16,17,18,19. This deficiency is particularly acute for complex tissues, such as the liver20,21, heart22,23, as well as stem cell and cancer niches24, where recapitulating the intricate multi-cellular architecture and dynamic biochemical gradients at a physiologically relevant scale remains difficult. Additionally, unlike bioprinting, which can fabricating large-scale, volumetrically complex biomimetic structures, organ-on-a-chip devices are often restricted to planar, micro-scale geometries25,26. Consequently, while organ-on-a-chip represent an advancement, they are often insufficient for fully simulating macroscale organ functions or generating the deep, volumetric tissue constructs required for patient-specific disease modeling27.
Three-dimensional (3D) printing and bioprinting have emerged as advanced technologies to overcome these architectural and material challenges. By enabling the fabrication of biomimetic models with precise control over cellular and matrix organization, 3D bioprinting is positioned to generate high-fidelity, systemic models25,28. These engineered tissues offer the structural complexity necessary to improve early-stage decision-making and de-risk the transition to human clinical trials.
Distinct from general reviews of NAMs5,29,30, this article emphasizes the pivotal role of 3D bioprinting technology specifically for fabricating in vitro systemic models tailored for drug screening, toxicology assessment, and disease modeling. Furthermore, unlike broad reviews of 3D bioprinting31,32,33, this paper maps current bioprinted in vitro models into a specific evaluation framework, emphasizing the critical role of 3D bioprinting in establishing complex in vitro pre-clinical models. This approach allows for a critical evaluation of the potential of current bioprinting technologies as standardized NAMs fabrication tools while rigorously assessing their limitations.
In this review, we provide a comprehensive analysis of leading 3D bioprinting technologies, including inkjet, extrusion-based, and vat photopolymerization, detailing their core mechanisms and recent applications in fabricating in vitro preclinical models. The discussion concludes by addressing current challenges, such as the evolving regulatory frameworks for 3D bioprinting-assisted NAMs, the technical hurdles in achieving multi-scale vascularization, and the necessity of maintaining anatomical fidelity across varying dimensions. Finally, it presents a forward-looking perspective on integrating biofabrication with data-driven design to forge predictive crosstalk between engineered tissues, ultimately driving therapeutic innovation.
Role of 3D bioprinting as a NAMs
The FDA Modernization Act 3.0 explicitly authorizes advanced NAMs for preclinical testing, with three-dimensional (3D) printing/bioprinting emerging as a notably powerful tool. The workflow28 of 3D printing typically commences with detailed imaging of the target biological structure via computed tomography (CT), magnetic resonance imaging (MRI), or X-ray, establishing precise spatial geometries. Subsequently, digital designs are generated based on guiding principles such as biomimicry or autonomous self-assembly, informing the spatial arrangement of cellular and structural elements. Selection of suitable biomaterials, including natural polymers, synthetic polymers, or decellularized extracellular matrices (ECM), is based on their biocompatibility and mechanical properties. Appropriate cell populations are then isolated and integrated with these biomaterials to formulate bioinks. These bioinks are precisely deposited into 3D constructions using inkjet-based, extrusion-based, or light-based bioprinting techniques. Following the bioprinting process, printed constructs undergo maturation within specialized bioreactors, promoting tissue differentiation, vascularization, and functional development, prior to their application either in vitro or as transplantable grafts (Fig. 2A)34,35. By effectively replicating the human tissue microenvironment, bioprinted models address critical limitations inherent to conventional animal models, often insufficient in predicting human-specific physiological and pathological responses, and traditional 2D cell culture systems, which lack the necessary three-dimensional context27.
A Typical process of 3D bioprinting. B Targets of 3D bioprinting in the current drug development pipeline. The figure was organized in Adobe Illustrator; icons are borrowed from BioRender.com.
Targets of 3D printed models in drug development
In alignment with the FDA drug development pipeline36,37, the principal utility of 3D printed models is currently concentrated in the early discovery and preclinical phases (Fig. 2B). During the discovery phase, the fabrication of patient- or disease-specific constructs, such as bioprinted tumor assembloids, facilitates high-throughput screening for target identification and lead compound validation37. By incorporating biologically relevant 3D microenvironments, this approach mitigates attrition risk early in the development cycle, effectively bridging the “valley of death” where candidate compounds frequently fail due to the poor translational fidelity of traditional 2D cultures and animal models5.
In the subsequent preclinical stage, bioprinted tissue constructs offer advantages for toxicology and pharmacodynamic assessments. Organotypic models, including lung38, skin39, and liver constructs40, demonstrate superior predictive capacity for drug screening of human-specific diseases, which are often obscured in animal studies. Under the framework of the FDA Modernization Act 2.0 (FDAMA 2.0)8, validated data derived from these bioprinted models can now be integrated into Investigational New Drug (IND) submissions. The contemporary regulatory landscape increasingly supports a hybrid testing strategy, wherein sponsors submit NAM data alongside conventional animal study results to substantiate the safety rationale for initiating human clinical trials.
Beyond preclinical safety, tissue chip technology is de-risking the clinical trial phase through the clinical trial on a chip paradigm15. This strategy prospectively models human population variability by utilizing 3D-printed microphysiological systems (MPS) populated with primary or stem-cell-derived cells from diverse donor cohorts41,42. By screening drug candidates across these stratified in vitro populations43,44, researchers can identify molecular signatures predictive of patient response. This data directly informs recruitment criteria and patient stratification, facilitating the discovery of novel biomarkers and more reliable clinical endpoints.
While traditional animal studies remain essential for specific systemic evaluations, the immediate advantage of 3D bioprinting lies in its capacity to enhance early-stage decision-making, preventing unsafe or ineffective candidates from progressing to costly late-stage trials. As validation protocols mature, bioprinted human tissues6,28 are poised to progressively replace specific animal-based assays, fulfilling the FDAMA 2.0 directive to advance preclinical methodologies toward more humane and scientifically robust outcomes. However, the regulatory acceptance of these advanced models requires a fundamental shift in validation strategies, as detailed in the following section5.
Validation of 3D printed models as NAMs
A shift is underway in how alternative toxicological methods are validated. This shift is highlighted by 2024 ICCVAM report, “Validation, Qualification, and Regulatory Acceptance of New Approach Methodologies”45. It explicitly recognizes that traditional validation processes, such as multi-year trials and direct comparisons to animal data, are often too slow for the rapid innovation seen in 3D tissue engineering45. Instead, the framework proposes a flexible, “fit-for-purpose” approach tailored to the specific application of the NAM”45.
The context of use (COU) is the central concept that defines the specific purpose of a NAM45. It requires that validation strategies align strictly with the intended application. Defining a precise COU is the first step in building scientific confidence and dictates all subsequent validation parameters5,45. Whether a model is designed for high-throughput screening or quantitative risk assessment, it must meet specific regulatory requirements5. For example, the FDA’s Center for Drug Evaluation and Research (CDER) may accept a bioprinted liver model for evaluating drug-induced liver injury (DILI), provided the method effectively detects known hepatotoxins5. Therefore, early communication between developers and regulators is essential to ensure study designs fit the regulatory landscape5.
Biological relevance is established by linking the NAM test system to an Adverse Outcome Pathway (AOP)45. An AOP is a framework that connects molecular events (such as receptor binding) to adverse health outcomes (such as organ failure)5. Mechanistic relevance has already been proven in several areas: skin sensitization assays follow Organization for Economic Co-operation and Development (OECD) Guideline 4975,29 and endocrine screening uses pathway models for receptor activity5. Additionally, validated vascular models can replicate endothelial behavior under physiological flow29. To further build confidence, validation studies should use biological reference compounds, substances with known effects in humans, as supported by updated FDA guidance on reproductive toxicity5,29.
Technical characterization ensures that the 3D bioprinted model is robust, reliable, and reproducible. This process involves a rigorous assessment of variability, ranging from bioink stability to printing parameters45. The ICCVAM framework recommends using industrial quality control tools to guide this process45. These include flowcharts to identify critical steps, cause-and-effect analyses to isolate variables like temperature changes, and control charts to monitor equipment45.
Finally, data integrity is the foundation of regulatory acceptance. Developers must evaluate their data pipelines and adhere to Good Laboratory Practice (GLP) principles before independent peer review45. The ICCVAM framework prioritizes transparency, specifically the public sharing of protocols and raw data, to build trust and speed up decisions45. Ideally, validation should undergo independent review by bodies such as NICEATM (U.S.), EURL ECVAM (EU), or JaCVAM (Japan)45. Ultimately, international harmonization is achieved through the OECD Mutual Acceptance of Data (MAD) agreement45. This ensures that safety data generated under OECD guidelines is accepted by all member countries, eliminating redundant animal testing and creating a global environment for NAMs45.
3D-bioprinted in vitro-complex models
The fabrication of in vitro models through 3D bioprinting has emerged as a unique strategy in drug testing and disease modeling, overcoming the limitations of traditional 2D cultures by replicating the intricate architecture and function of native human tissue28,46. This review focuses on the three principal bioprinting modalities (jetting-based, extrusion-based, and vat photopolymerization), analyzing their distinct capabilities in fabricating functional, systematic tissue models. Unlike static 2D assays, these biofabricated constructs serve as high-fidelity systematic models, defined here by their capacity to support architectural heterogeneity (the precise spatial arrangement of multiple cell types) and functional perfusion networks essential for nutrient exchange and hemodynamic simulation (Fig. 3).
This diagram illustrates the model systems used to recapitulate the hierarchical organization of the human body. As the biological scale increases from subcellular components to systemic levels, the complexity of the modeling modalities rises correspondingly. The figure further highlights common bioprinting methodologies employed to fabricate systematic in vitro models. The figure was organized in Adobe Illustrator; icons are borrowed from BioRender.com.
In the following sections, we evaluate recent advancements in these commonly used modalities through a multi-parametric assessment framework. This framework investigates the potential of these printed structures to serve as in vitro systematic models and NAMs. First, we examine the biological relevance, specifically focusing on the integration of human-derived cells to mitigate species-specific discrepancies found in animal models. Second, we assess the complexity of the model, analyzing how each printing technique achieves structural fidelity and whether they support dynamic perfusion. Finally, we categorize these models by their target endpoints (e.g., specific disease phenotypes and applications such as drug screening or disease modeling) and critically review the validation methods employed. This ensures that the established models are not only structurally sophisticated but also functionally validated against physiological benchmarks to serve as effective tools for regulatory decision-making.
Jetting-based bioprinting
Jetting-based bioprinting, especially the inkjet bioprinting, is an extensively investigated non-contact additive manufacturing modality for depositing cells and biomaterials. Its operation relies on on-demand droplet ejection, typically via thermal, piezoelectric, or electrostatic actuation, to deliver picoliter-to-nanoliter bioink droplets with spatial precision and low mechanical insult (Fig. 4A)47. Key features of this strategy are summarized in Table 1. A fundamental challenge in this domain is the definition of printability, which is governed by a rigorous balance of inertial, viscous, and surface tension forces48. This balance is quantified by the dimensionless Z number, defined mathematically as the inverse of the Ohnesorge number (Oh-1)48. The Z number encapsulates the fluid’s physical state by relating the Reynolds number (Re) to the Weber number (We) through the equation (Z=1/Oh={text{Re}}/sqrt{We}=sqrt{rho cdot gamma {L}_{c}}/eta), where ρ is density, γ is surface tension, Lc is the characteristic length (nozzle radius), and η is viscosity. For stable drop-on-demand (DOD) printing, the Z number must theoretically fall within a specific window, typically between 1 and 10, though some experimental setups tolerate values up to 1449. Deviations from this range result in distinct failure modes. Values below 1 lead to viscous dissipation, where the fluid cannot overcome surface tension to eject, while values exceeding the upper limit result in satellite formation because excess inertia causes the liquid ligament to fragment into secondary droplets49.
A Jetting-based bioprinting. B Extrusion-based bioprinting, incluidng direct ink writing and embedded ink writing. C Two typical vat photopolymerization mehods: two-photon polymerization and digital light processing. D Advanced vat photopolymerization method-volumetric bioprinting, including light-based, acoustic-based, and hybrid strategies. The figure was organized in Adobe Illustrator; icons are borrowed from BioRender.com.
Beyond fluid physics, the optimization of inkjet bioprinting requires a careful analysis of factors affecting cell viability, which generally remains high at 80-95% despite the mechanical stresses involved. Shear stress is widely recognized as the most critical factor and is primarily caused by the sharp velocity gradients generated as the fluid is forced through the micro-nozzle at high speeds50. Current results indicate that shear stresses exceeding 10 kPa significantly compromise long-term cell proliferation and can trigger apoptotic pathways50. Besides, recent studies also indicate that droplet volume plays a critical role in viability due to evaporation kinetics. A minimum droplet volume of 20 nL per spot is required to mitigate evaporation-induced cell damage and maintain high viability during the printing process51. Thermal effects also present distinct challenges, particularly in thermal inkjet systems where the heating element can reach temperatures exceeding 300 °C. However, due to the microsecond duration of the heating pulse, the bulk temperature of the bio-ink rises only 4–10 °C, minimizing damage to the cellular payload47. Moreover, during the solidification stage of bioprinting, various crosslinking methodologies, including light52, chemical53, ionic54, acoustic55, and enzymatic56 mechanisms, can induce significant alterations in transcriptomic dynamics, thereby shifting cellular behavior57. While these phenotypic and transcriptional responses to physicochemical cues are well-established in general cell biology58,59,60,61,62, they remain insufficiently characterized within the specific context of bioprinting63. Consequently, further research is urgently needed to systematically evaluate the impact of these diverse printing and consolidation modalities on both short-term stress responses and long-term cellular function.
Category of Inkjet bioprinting methodologies is defined by distinct actuation mechanisms that dictate the balance between resolution and material capabilities, critical features are summarized in Table 1. Standard inkjet systems, utilizing nozzles with diameters of 20–100 µm, achieve resolutions in the range of 20–50 µm64. While thermal inkjet (TIJ) systems are cost-effective, piezoelectric inkjet (PIJ) technology offers superior volumetric control without thermal stressors. A third category, electrohydrodynamic (EHD) jetting, utilizes high-voltage electric fields to achieve significantly higher resolutions of 2–5 µm64. Despite these capabilities, the technology faces specific material constraints. Standard inkjet systems are generally limited to low-viscosity fluids, typically under 20–30 mPa·s64, which restricts the use of mechanically dense hydrogels necessary for structural support.
Despite these material constraints, bio-applications of this technology have broadened markedly over the past decade, with critical features emerging for selected cases of fabricating in vitro biological models (Table 2). Specifically, inkjet systems have been used to fabricate biomimetic constructs at high throughput, including vascular-like patterns and multicellular architectures relevant to regenerative medicine38. Their compatibility with miniaturized drug-screening platforms is well documented, allowing patient-derived cells to be arrayed in small tissue units for efficacy testing under disease-relevant conditions38. Beyond organoid arrays, sequential droplet deposition of cell-laden hydrogels supports the assembly of multilayered, 3D tissue-like constructs with promise for disease modeling and target discovery65. Applications also extend to microphysiological systems, where printed cell arrays recapitulate organ-level behaviors in liver and cardiac models66. Most recently, this capability has been adapted to pulmonary pathology, with all-inkjet-printed 3D alveolar barrier models proving effective for mimicking fibrosis and evaluating anti-fibrotic therapeutics67.
Extrusion-based bioprinting
Extrusion-based bioprinting is the main modality for building centimeter-scale, cell-laden constructs. In this process, a continuous filament of bioink is extruded through a nozzle and patterned along programmed toolpaths to assemble 3D architectures. Pneumatic or mechanically driven systems (plunger/screw) accommodate viscous hydrogel inks and pastes, including composites with microfibers or particulates, while supporting high cell loadings within each strand. Compared with droplet methods, extrusion tolerates a broader rheological window and is better suited for depositing bulk volumes with embedded cells, making it a workhorse for tissue- and organ-scale fabrication. In this section, we focus on two widely used strategies: (i) traditional extrusion into air, where filaments rapidly solidify to support subsequent layers; and (ii) fluid bath-assisted extrusion (e.g., embedded/support-bath printing), which provides in situ support to expand printable materials and geometries. Key features of these strategies are summarized in Table 1.
Direct ink writing (DIW)
Direct ink writing (DIW), also termed direct-write assembly, is an extrusion-based modality in which viscoelastic inks are dispensed through fine nozzles and patterned along programmed toolpaths to form continuous filaments that assemble three-dimensional architectures with specified composition and topology (Fig. 4B)68,69. To determine good printability of DIW, researchers utilize the printability number (Pr)70 which is a quantitative metric derived from the geometry of pores in a printed lattice. It is calculated as the ratio of the actual pore circularity to the theoretical square shape using the equation (Pr ={L}^{2}/16A), where L is the perimeter of the pore and A is the area. The value of Pr = 1 indicates perfect fidelity. Deviations indicate failure modes where values < 1 suggest under-gelation leading to filament spreading and values > 1 indicate over-gelation or irregular filament formation70. Additionally, for printable inks, the material must exhibit a gel-sol transition under high shear strain and a rapid sol-gel recovery upon cessation of shear. Ideally, the ink should show storage modulus (G’) > loss modulus (G”) at rest (solid-like behavior) to ensure particle suspension and shape retention68. Under the high shear of dispensing, the network structure breaks down, and G” may approach or exceed G’, facilitating flow. The rate at which G’ recovers after deposition determines the thixotropic time of the ink68. A rapid recovery is essential to prevent filament fusion and ensure that the printed structure does not deform under its own weight68,71.
The incorporation of living cells introduces biological constraints that often conflict with these rheological ideals. The primary factor affecting cell viability is shear stress ((tau)) which is generated by the velocity gradient within the nozzle. For a fluid flowing through a cylindrical channel, the shear stress varies linearly with the radial position (r) according to the equation72. (tau (r)=rcdot varDelta P/2L) where ΔP is the pressure drop and L is the nozzle length. Empirical studies have established distinct thresholds where shear stresses < 5 kPa define a “safety zone” with high viability (>90%) while stresses > 14 kPa fall into the “lethal zone” that triggers immediate lysis or irreversible apoptotic cascades72. To mitigate these forces, conical nozzles are preferred over cylindrical needles as they eliminate abrupt contractions and reduce the residence time of cells in high-stress regions72. Furthermore, matrix crosslinking strategies significantly impact viability by altering the processing window68,71. By employing post-printing crosslinking, researchers can extrude at lower pressures to reduce the shear stress experienced by the cells while still achieving the final structural rigidity necessary for tissue formation68,71.
Leveraging this foundation, recent studies demonstrate DIW’s utility for pharmacology-ready, perfusable models and anatomically faithful soft-tissue constructs (Table 3). For instance, a sacrificial-free DIW (SF-DIW) strategy produced one-step hollow networks without sacrificial templates40, utilizing a bioink composed of gelatin, alginate, and GelMA pre-crosslinked with CaSO4 to ensure immediate structural fidelity. This approach enabled the integration of micropumps and an agar casting step for physical encapsulation, creating a robust perfusion system for liver metastasis models populated with HepaRG hepatocytes and human umbilical vein endothelial cells (HUVECs)-lined vascular channels. Functional assessment under drug challenge revealed that the perfused hepatic environment successfully upregulated albumin and hepatocyte nuclearfactor4alpha (HNF4α) expression (confirmed via qRT-PCR and Western blot) and bioactivated the prodrug ifosfamide to inhibit the growth of co-cultured breast cancer (MCF-7) spheroids. Complementarily, a sacrificial-scaffold-assisted (SSA) DIW route created aortic valve prostheses with overhanging leaflets and sinus geometry73. In this study, a thermosensitive Pluronic F127 fugitive lattice supported the deposition of alginate/gelatin/Laponite nanoclay bioinks, where the nanoclay induced a “house-of-cards” microstructure via electrostatic interactions to enhance yield stress. The gentle removal of the sacrificial support by lowering the ambient temperature preserved the intricate internal cavities and one-way valvular function, validated through fluid-structure interaction simulations and mechanical cycling, while maintaining viability for mouse and human cell types. At the bioprinting–bioelectronics interface, DIW now patterns conducting, semiconducting, dielectric, and hydrogel phases within a single build, yielding conformal sensors and electrodes that provide continuous readouts (strain, impedance, electrophysiology) for organoids and tissue-chips during compound exposure74. Additionally, DIW has also enabled disease-relevant microtissues with precisely tuned extracellular matrices to dissect tumor, ECM interactions, and invasion dynamics, offering a controllable platform that can be coupled to perfusion and drug dosing75.
Despite these advances, the technology is constrained by inherent trade-offs between resolution, speed, and cell density. The resolution of DIW is physically bounded by the nozzle diameter, generally limiting features to filaments greater than 100 µm, which is significantly coarser than the 20–50 µm resolution achievable with inkjet systems76. Pushing below this limit exponentially increases the shear stress and the risk of nozzle clogging, particularly when printing high-density cell slurries76. Furthermore, the process exhibits an inverse relationship between resolution and throughput, reducing nozzle size to improve detail necessitates a drastic reduction in flow rate, making the fabrication of large-scale organ constructs prohibitively slow and potentially exceeding the viability window of the encapsulated cells76. Finally, the layer-by-layer deposition creates a “staircase” effect on curved surfaces, which can result in surface roughness that influences cell attachment and complicates the fabrication of anatomically precise geometries like heart valves71.
Embedded ink writing (EIW)
Embedded ink writing (EIW), also termed embedded 3D printing or support-bath–assisted printing, is a material-extrusion strategy in which liquid inks are deposited within a viscoelastic support medium rather than in open air77,78,79. In EIW, the bath (typically a yield-stress hydrogel or granular gel) behaves as a Bingham-type material80: it fluidizes locally under the shear field of the translating nozzle and rapidly re-solidifies once the nozzle passes, thereby immobilizing the extruded filament in three dimensions (Fig. 4B). Ideally, this rheological behavior is described by the Herschel-Bulkley model ((sigma ={sigma }_{y}+K{dot{gamma }}^{n}))80, where σ is the shear stress, representing the total force the bath exerts on the nozzle. The yield stress (σy) represents the critical threshold the material must overcome to flow. Below this value, the bath acts as a solid to trap the ink. The Consistency Index (K) represents the viscosity or thickness of the fluid once it begins flowing. The shear rate ((dot{gamma })) corresponds to the speed of deformation (nozzle speed), and the flow index (n < 1) characterizes the shear-thinning behavior that reduces viscosity as the nozzle moves faster80. Furthermore, the spatial precision of the print is governed by the Oldroyd Number, a high Od ensures the fluidized region is tightly confined to the nozzle tip, preventing the disturbance of adjacent layers77.
This self-healing, shear-thinning response provides omnidirectional mechanical support throughout printing, suppressing sag due to gravity or surface tension77. Specifically, to maintain geometric fidelity, the support bath must satisfy two fundamental stability inequalities: the yield stress must exceed the buoyant stress exerted by density mismatches ((varDelta rho Vg{Y}_{crit} < {sigma }_{y}A)) to prevent sedimentation, where (varDelta rho) is the density difference between the ink and bath (which should ideally be zero), V is the volume of the printed feature, A is the surface area of the extruded ink, and Ycrit is the critical vertical displacement or failure distance the object sinks before the print is ruined77. Second, it must structurally resist interfacial tension ((R{sigma }_{y}/gamma > 1)) to prevent the filament from breaking up into droplets due to the Plateau-Rayleigh instability. In this ratio, R is the nozzle/filament radius and γ is the interfacial tension between the ink and the bath. A result greater than 1 indicates that the bath’s solid-like strength is sufficient to overpower the surface tension forces that try to bead the filament into spheres77. Following deposition, the ink, often a hydrogel precursor or cell-laden bioink, is crosslinked in situ (thermal gelation, enzymatic or ionic cross-linking, photopolymerization) while still embedded, fixing its shape. The support matrix is then removed gently to release a self-supporting construct. By obviating sacrificial scaffolds or layer-by-layer supports, EIW enables freeform fabrication from soft biomaterials with high fidelity. Filament diameters on the order of 100–200 µm are routinely achieved with smooth, continuous deposition in three dimensions80. Notably, the bath itself can be functionalized, for example, loaded with diffusing cross-linkers that initiate interfacial gelation, to promote gentle consolidation of printed features81,82. In aggregate, the mechanism markedly expands the printable material space, including delicate hydrogels and living bioinks, by providing a biocompatible milieu in which intricate architectures can be “written” and stabilized in situ.
The FRESH (freeform reversible embedding of suspended hydrogels) approach exemplifies early EIW capabilities83,84. Using a thermoreversible gelatin microgel bath, FRESH enabled 3D printing of soft inks such as alginate, collagen, and fibrin with unprecedented geometric freedom (Table 3)83. Complex anatomical architectures, including branched coronary networks, trabeculated embryonic heart models, gyroidal brain structures, and even an adult femur replica, were produced at ~200 µm resolution by extruding hydrogels into the gelatin slurry and crosslinking in situ (e.g., via ionic or pH triggers)83. After warming, the gelatin support melted away, yielding robust constructs that faithfully captured internal and external organ geometries. These results further showed that even very soft hydrogels (elastic modulus < 500 kPa) can be additively manufactured into complex forms when printed inside a supportive matrix83. As a concrete demonstration of translational potential, Mirdamadi et al.85 used FRESH to fabricate a full-scale alginate human heart model derived from patient MRI that mimicked cardiac tissue mechanics; the life-sized heart could be sutured and handled like a native organ, offering a realistic platform for pre-operative rehearsal and device testing.
Beyond acellular scaffolds, embedded printing has been extended to living inks with high viability and spatial fidelity. Several critical factors impact cell viability in this process: (1) Shear stress, which is significantly mitigated in EIW because the support bath allows for the use of low-viscosity inks (1–100 Pa·s) that reduce shear forces on cell membranes77; (2) Extrusion pressure, where lower driving pressures prevent physical compression and depressurization shock at the nozzle exit80; and (3) Bath biocompatibility, which requires careful management of osmolarity to prevent osmotic shock and cell lysis77,80. Bhattacharjee et al. showed that a granular support (jammed microgels) permits printing not only of polymers and colloids but also of living cells as the ink itself86. Using a yield-stress Carbopol microgel, the research team wrote thin-walled shells, knotted loops, and branching tubular networks from silicones, hydrogels, and aggregates of cells. Printed structures remained stably embedded for hours. Polymeric inks were subsequently crosslinked, while cellular features were cultured directly within the bath. Calcein staining confirmed high viability of human mammary cell spheroids for at least five days post-printing, indicating minimal cytotoxic burden from embedding and write-induced fluidization86.
A landmark in tissue engineering came from Kolesky et al.87, who co-printed vascularized, heterogeneous cell-laden tissues within a gelatin-based support bath (Table 3). In this study, multiple bioinks were embedded and printed together to form a ~ 1 cm-thick construct containing interpenetrating vascular channels and encapsulated cell populations (e.g., fibroblasts, epithelial cells) in an ECM hydrogel. A fugitive Pluronic F127 ink defined a vascular network inside GelMA. Following matrix crosslinking, the sacrificial phase was evacuated, leaving perfusable microchannels that were endothelialized to mimic blood vessels. These densely cellular tissues were sustained by convective nutrient delivery and maintained in vitro over extended culture, with high bulk viability (~90% post-printing reported in similar systems87) enhanced by perfusion.
Additionally, materials engineering of the support phase has broadened EIW’s reach of printing standalone vascular architectures. Compaan et al.81 introduced a gellan-gum fluid gel, a yield-stress bath with thermal/pH stability that can be doped with gentle cross-linkers (e.g., Ca2+, enzymes) to diffuse into printed strands and cure sensitive precursors (fibrinogen, alginate) in place. Cell-laden constructs printed in gellan exhibited excellent viability (often > 80–90% post-printing) and high fidelity, with the aqueous matrix maintaining hydration during printing. In parallel, Patrício et al.88 formulated a xanthan-gum (XG) bath as a continuous pseudoplastic gel, enabling rapid freeform printing with out-of-plane features and effective up-scaling to clinically relevant dimensions. Acting as a photocurable reservoir, XG supported on-the-fly UV crosslinking of cell-laden GelMA (gelatin methacryloyl), yielding perfusable, cell-populated constructs with embedded channels proposed as tunable in-vitro models (e.g., cardiac, tumor) and potential organ-on-chip platforms given their capacity for long-term viability and perfusion.
Personalized, clinically relevant models underscore EIW’s integrative power. Noor et al.89 bioprinted autologous cardiac patches and a miniature heart using patient-derived cells and ECM-based bioinks (Table 3). From a small omental biopsy they generated (i) induced pluripotent stem cells that were differentiated into cardiomyocytes and endothelial cells, and (ii) a decellularized patient-specific ECM hydrogel serving as the bioink. Using embedded printing, they produced perfusable, anatomically matched cardiac patches exhibiting organized tissue structure and synchronous contractions. As a result, viability remained high, and endothelialized channels supported nutrient delivery through the thickness. Subsequently, a small-scale cellularized human heart with patient-specific geometry was also printed as proof of concept. While not fully functional, these constructs suggest immediate use in patient-specific drug screening and, longer-term, in autologous repair.
SWIFT (sacrificial writing into functional tissue) further addresses organ-level cell density and vascularization. Skylar-Scott et al.90 packed hundreds of thousands of stem-cell-derived organoids (organ building blocks, OBBs) into a dense, ~200 million cells mL−1 living matrix and embedded-printed a sacrificial gelatin lattice directly through the tissue. Upon warming to 37 °C, the gelatin was removed to leave pervasive channels that were immediately perfused. Perfused tissues remained fully viable, whereas equally sized non-perfused controls developed rapid core necrosis. In cardiac matrices, discrete cardiomyocyte clusters fused over a week into synchronously beating tissue with >20-fold stronger contractions, approaching native-like function. By combining OBBs with embedded vascular writing, SWIFT provides an integrated route to centimeter-scale, high-density, perfusable tissues, valuable now for in-vitro pharmacology and development studies, and foundational for eventual organ-scale biomanufacturing.
In summary, EIW leverages yield-stress supports, rapid self-healing, and in-bath curing to print complex, cell-rich architectures with high fidelity while preserving viability. The approach enables multi-material and multi-cellular integration and the formation of perfusable networks, spanning applications from anatomical scaffolds and organ-on-chip systems to thick, vascularized tissues and mini-organs for drug testing and disease modeling. As support-bath chemistries and printing methodologies advance, EIW is poised to assume a central role in tissue engineering and organ fabrication80,91.
Vat photopolymerization
The landscape of tissue engineering has been reshaped by the advance of vat photopolymerization technologies that offer an alternative to the shear stress limitations of nozzle-based extrusion. Within this domain three distinct modalities, including two-photon polymerization (TPP), digital light processing (DLP), and volumetric bioprinting (VBP) occupy orthogonal niches defined by their optical mechanisms and biological capabilities. The primary divergence lies in their excitation strategies, where TPP leverages non-linear optical phenomena using near-infrared (NIR) femtosecond lasers to achieve sub-micron resolution below the diffraction limit of light92. Conversely, DLP functions as a layer-by-layer modality projecting dynamic photomasks typically using light wavelength from 365 to 980 nm93,94. VBP employs tomographic back-projection to solidify an entire 3D volume simultaneously by projecting cumulative dose patterns into a rotating vial using 375 nm or visible light (450–700 nm)95,96.
These distinct optical mechanisms necessitate specialized ink formulations to ensure fabrication fidelity. For visible light systems like DLP and VBP the chemical Lithium phenyl-2,4,6-trimethylbenzoylphosphinate (LAP) has emerged as the standard photoinitiator due to its high water solubility and cytocompatibility93,95. However, the role of additives varies critically as DLP relies on synthetic photoabsorbers such as Tartrazine to limit light penetration depth according to the Beer-Lambert law, which prevents cure-through and enhances Z-axis resolution94,97. On the other hand VBP demands optical transparency often requiring refractive index matching agents like Iodixanol to minimize cell-induced light scattering that would otherwise distort the projected tomograms96. TPP formulations are even more specialized often utilizing efficient but potentially less biocompatible initiators such as P2CK or IP-Dip resins98 to maximize the two-photon absorption cross-section which necessitates rigorous washing steps post-fabrication.
Ultimately, the choice of technology is governed by the specific biological constraints of the application. While VBP offers the highest cell viability profile by printing centimeter-scale organs in seconds which eliminates shear stress and time-dependent cell death it requires high cell densities such as 50 × 106 cells/mL96 that complicate light propagation. In contrast, DLP faces challenges with cell sedimentation during longer print times and potential oxidative stress from free radicals though the shift to 405 nm has improved outcomes compared to UV-based predecessors93,94,99. Despite operating within the safe optical therapeutic window of NIR light TPP is severely limited by fabrication speed because the point-by-point scanning process is prohibitively slow for large constructs potentially leaving cells outside optimal culture conditions for extended periods although it remains unmatched for defining sub-cellular topography92,100,101.
Two-photon polymerization (TPP)
TPP is a laser microfabrication strategy in which a femtosecond near-IR focus drives polymerization only within a sub-femtoliter voxel via nonlinear two-photon absorption. Because curing is confined to the focal volume, intricate 3D features, including capillary-scale conduits (~10–30 µm), can be “written” inside photosensitive hydrogels without affecting surrounding material102 (Fig. 4C). Cells can be present in the resin yet remain largely unexposed outside the focal spot, supporting direct encapsulation. Critical features are summarized in Table 1.
Recent advances have pushed TPP toward practical bioprinting speeds through highly reactive, cytocompatible chemistries. A gelatin-norbornene (Gel-NB) thiol-ene system achieved scan speeds up to 1000 mm s⁻¹ while maintaining cell survival. Meanwhile, stiffness could be tuned (~0.2–0.7 kPa) by laser parameters, and encapsulated cells spread, migrated, and proliferated over multi-week culture103. Exploiting this capacity for architectural heterogeneity and stiffness control (Table 4), Sharaf et al.104 fabricated 2.5D nano-pillar arrays and 3D micro-cages to guide primary rhesus macaque microglia toward a homeostatic state. By presenting these immune cells with specific topographical cues, the authors fostered a ramified resting phenotype that is difficult to maintain in vitro, validating the model through quantitative morphological analysis of branch complexity and volumetric cell colonization. Using TPP, perfusable microvascular networks with 10–30 µm channels have been patterned within cell-laden gels, yielding endothelialized, stable microvessels suitable for organ-on-chip applications102. Expanding on this to create multi-tissue interfaces (Table 4), Mandt et al.105 utilized TPP to fabricate a biomimetic placental barrier directly within a microfluidic device using a gelatin-based hydrogel (GelMOD-AEMA). This high-resolution villous membrane separated a fetal compartment lined with HUVECs from a maternal compartment of trophoblasts, creating a functional barrier model validated via time-dependent glucose transport assays. Beyond photopolymerization, ultrafast lasers can also sculpt vasculature in native matrices: femtosecond cavitation molding opened 20-60 µm microchannels inside living collagen gels that were subsequently endothelialized, enabling vascularized tumor models with sustained perfusion106.
Digital light processing (DLP)
Digital light processing (DLP) is one of vat photopolymerization bioprinting methods in which a digital micromirror device (DMD) projects patterned light to selectively solidify photosensitive bioinks layer by layer93,107,108,109. Unlike laser-based stereolithography (SLA), which traces a single focal point, DLP cures an entire cross-section in one exposure, markedly accelerating print speed93,107,108,109,110. Masks derived from computer-aided designs govern complex geometries. After each exposure the build platform translates to refresh resin, and the next pattern is projected. This layer-wise projection enables intricate three-dimensional hydrogel constructs with microscale fidelity, where resolution is set primarily by pixel pitch and projection optics, typically on the order of tens of micrometers (~25–50 μm)93 (Fig. 4C). Key characteristics are summarized in Table 1. Consequently, DLP can reproduce fine cellular architectures that are difficult for extrusion systems limited by ~200–500 μm nozzles93, while the non-contact curing yields smooth surfaces and avoids mechanical shear on cells.
DLP bioprinting has been applied across engineered scaffolds and multicellular disease models (Table 4). Xu et al.111 introduced a two-step DLP workflow to fabricate polyacrylamide–alginate hydrogels with tunable stiffness for cardiac and vascular applications. Complex three-dimensional architectures were first patterned by DLP, followed by secondary ionic crosslinking via Fe3+ soaking to adjust modulus across 15.8–345 kPa111. The process achieved ~10 μm feature precision and enabled matching of diverse soft-tissue mechanics. Cardiac muscle scaffolds supported organized, synchronously beating myocardium in vitro, whereas vascular patterns promoted branching endothelial networks, demonstrating that the fidelity of printed microarchitecture, together with appropriate stiffness cues, drives native-like morphology and function and provides platforms for maturation studies and cardiotoxicity testing111.
For hepatic tissue modeling, Ma et al.112 leveraged DLP to create deterministically patterned human liver constructs for drug screening. A three-dimensional hexagonal, lobule-like hydrogel was printed with human induced-pluripotent-stem-cell-derived hepatic progenitor cells (hiPSC-HPCs), together with supportive endothelial and mesenchymal cells, to recreate the native tri-cellular microenvironment. Using a customized DLP system, physiologically sized hexagonal units were fabricated within seconds under minimal UV exposure. During weeks of perfused culture, the tri-culture exhibited enhanced polarity and organization, elevated expression of liver-specific genes, increased albumin and urea secretion, and inducible cytochrome P450 activity comparable to adult hepatocytes. Notably, rifampicin triggered robust CYP3A4 upregulation, indicating functional drug metabolism112. These data underscore DLP’s strength in rapidly fabricating high-fidelity, multicellular liver models as patient-specific alternatives to animal testing for early-stage screening and disease modeling112.
DLP’s optics-defined patterning has also enabled in vitro disease models that recapitulate pathological niches. Zhong et al.113 reported a bioprinted pterygium construct comprising primary human conjunctival stem cells (hCjSCs), immune cells, and vascular endothelial cells. Projection printing maintained high viability and phenotypic integrity of the delicate hCjSCs. Within the three-dimensional hydrogel, embedded cells exhibited transcriptomic signatures of inflammation, angiogenesis, and epithelial-mesenchymal transition that closely matched clinical pterygium tissue. As the first three-dimensional in vitro model of pterygium, this system enables mechanistic studies of pathogenesis and screening of anti-inflammatory or anti-angiogenic therapies on patient-specific constructs.
Oncology applications similarly benefit from DLP’s capacity to rapidly impose spatial heterogeneity. Tang et al.114 bioprinted a glioblastoma (GBM) model that captures the tumor’s biophysical complexity. A tri-regional construct, GBM cell-laden tumor zone, cell-free ECM-like zone, and endothelialized vascular region, was produced with region-specific stiffness using tunable gelatin-methacrylate bioinks. Mechanical cues dictated phenotype and therapy response: cells in stiffer, tumor-mimetic regions adopted a mesenchymal, invasive state with extensive vascular sprouting and heightened resistance to temozolomide, whereas cells in softer, brain-like regions retained a more proliferative, classical phenotype. These region-dependent behaviors mirror patient tumors and can be replicated with patient-derived cells, highlighting DLP as a platform for mechanistic studies under realistic biophysical conditions and for screening therapies that must penetrate or modulate tumor mechanics.
Collectively, DLP bioprinting unites optics-limited, high-resolution patterning with gentle, rapid fabrication to yield living constructs that emulate native microarchitecture and function. Demonstrated outcomes span synchronously beating myocardium and vascular networks111, organotypic liver units with adult-like metabolic capacity112, and complex disease niches such as ocular fibrosis113 and GBM114. Continued advances in biocompatible photoinitiators, multi-material projection110,115, and expanded build volumes will further extend DLP’s impact, enabling increasingly realistic tissues-on-demand for drug screening, disease modeling, and regenerative medicine.
Volumetric bioprinting (VBP)
Volumetric bioprinting (VBP) refers to a series of techniques that form entire three-dimensional constructs in a single exposure or continuous operation rather than by layer-by-layer deposition. By curing or structuring the build volume directly, VBP enables fabrication in seconds to minutes of complex, cell-laden architectures without auxiliary supports, accommodating overhangs, enclosed cavities, and interwoven vascular networks with high geometric fidelity95,116,117,118. Recent developments report unprecedented print speeds alongside fine feature definition, yielding tissues that maintain high cell viability while preserving intended microarchitecture118. Conceptually, VBP modalities can be grouped by energy source: light-based approaches rely on photopolymerization within the bulk, acoustic approaches use focused ultrasound to pattern materials volumetrically, and hybrid schemes couple these fields to combine speed, resolution, and biocompatibility within a single process, positioning VBP as a complementary route to rapid, support-free biofabrication in this review. Additionally, critical features of this technology are summarized in Table 1.
Light-based VBP
Light-based volumetric bioprinting (VBP) relies on volumetric photopolymerization of a photocurable bioink to solidify an entire three-dimensional object simultaneously. A landmark implementation is computed axial lithography (CAL) (Fig. 4D), introduced by Kelly et al.119 CAL projects a sequence of two-dimensional light fields, computed by reverse tomography, into a rotating vial of photoresponsive resin. As the vial spins, the cumulative dose at each voxel crosses the polymerization threshold and the target geometry “appears” in bulk. This tomographic strategy eliminates layer-wise fabrication and external supports, enabling smooth surfaces, complex overhangs, and enclosed cavities. Centimeter-scale parts can be produced in a single, continuous exposure on the order of tens of seconds, with initial demonstrations achieving ~100 µm resolution and printing simple plastic and hydrogel forms in under a minute120. CAL’s material scope and resolution have since expanded. Notably, Toombs et al.121 adapted a microscale CAL process to fabricate intricate silica-glass microstructures with features of only a few tens of micrometers, underscoring that volumetric light-based printing can extend beyond soft polymers to inorganic systems.
Holographic projection has recently advanced light-based VBP by improving optical efficiency and feature control. Álvarez-Castaño et al.122 replaced amplitude-only CAL projections with phase-controlled holograms generated by a spatial light modulator, yielding ~20-fold higher light-delivery efficiency than standard patterned illumination. Using a low-power 40 mW laser, they printed millimeter-scale objects with ~30 µm features in under a minute, and, crucially, maintained high fidelity in scattering, cell-laden hydrogels (~5 × 105 cells mL⁻¹). Viable cells within the printed volumes demonstrated that holographic tomographic VBP can directly bioprint tissue constructs in soft bioinks despite multiple-scattering environments.
Progress in resin chemistry has been equally important for biocompatibility and function. Rizzo et al.123 formulated a “photoclick” bioresin based on gelatin-norbornene (GelNB) that undergoes rapid thiol-ene step-growth, enabling volumetric prints in ~10 s with near-100% immediate post-print viability. Low radical burden and fast gelation supported myoblast proliferation and differentiation into contractile myotubes within the printed constructs. By tuning the degree of functionalization, polymer content, and crosslinker concentration, hydrogel stiffness spanned from the tens of pascals to ~15 kPa, accommodating diverse tissue targets. Moreover, Lian et al.124 showed that pristine decellularized extracellular matrix (dECM) bioinks, heart and meniscus (Table 4), ~1% w/v, can be volumetrically printed in 30–60 s using a visible-light tris(2,2-bipyridyl)dichlororuthenium(II) hexahydrate/sodium persulfate (Ru/SPS) initiator without synthetic polymer additives. Because VBP decouples fabrication from load-bearing rheology, the intrinsic softness of dECM is no hindrance: centimeter-scale, anatomically inspired constructs (mini heart, ear, meniscus) with encapsulated cells were produced within tens of seconds. Cardiac dECM constructs seeded with cardiomyocytes exhibited robust spreading, sarcomeric maturation, and synchronous contractions, while meniscus dECM constructs with human mesenchymal stem cells underwent chondrogenic differentiation with cartilage-matrix deposition over time.
Tomographic VBP has also been extended to protein-based hydrogels and composites. Xie et al.125 demonstrated volumetric printing of unmodified silk protein bioinks (silk fibroin and sericin) achieving ~50–60 µm resolution. Dense cell encapsulation (5 × 106 cells mL⁻¹) yielded > 90% viability with sustained proliferation over 14 days. Distinct cell behaviors emerged across the two matrices: sericin supported spreading and growth, whereas stiffer fibroin constrained spreading. As a proof of concept toward implantable devices, the team printed a threaded bone-fixation screw from silk fibroin hydrogel, post-cured it, and seeded mesenchymal stem cells. Cells remained nearly 100% viable, proliferated, expressed osteogenic markers, and the screw fit stably in an ex vivo bone-defect model.
An alternative optical VBP strategy, Xolography, defines a curing zone by the intersection of two differently colored light fields. Regehly et al.126 introduced the concept in a dual-color photoinitiator (DCPI), where a sheet of light at λ1 (375 nm) photochemically “activates” a thin region and a second, dynamically patterned beam at λ2 (450–700 nm) induces polymerization only where it overlaps the activated sheet. Continuous motion of the optics or resin “draws” the object volumetrically with fully smooth surfaces and without discrete layers (Fig. 4D). In hard photopolymers, Xolography produced complex centimeter-scale objects (e.g., a boat inside a bottle) with ~25-50 µm features in under a minute. However, translation to bioprinting required aqueous, cytocompatible dual-color chemistry. Wolfel et al.127 realized “bioxolography” in GelMA using a modified initiator system augmented with diphenyliodonium chloride (DPI) and N-vinylpyrrolidone (NVP), enabling > 1 cm3 cell-laden constructs with ~20 µm positive-feature resolution (and ~125 µm for negative voids) within minutes (Table 4). The same platform supported multimaterial switching during a single build and engineered gradients via grayscale-controlled crosslink density to tailor local stiffness. Simultaneously, embedded fibroblasts and stem cells remained viable and supported long-term culture with matrix deposition. In parallel, Stoecker et al.128 demonstrated spatial control of mechanical properties by modulating beam overlap and exposure, encoding soft-to-stiff transitions that mimic native tissue heterogeneity. Using a compact, suitcase-scale printer derived from the Xolo “Xube” platform, they produced viable, cell-laden structures with ~20 µm features and uniform cell distributions, evidence that Xolography can be engineered into practical bioprinting systems.
In summary, tomographic methods such as CAL and multi-beam strategies such as Xolography furnish light-based VBP with unmatched speed and geometric freedom for complex, cell-rich architectures. With continued refinement in photochemistry, multimaterial control, and optical delivery, and with attention to scalability and cell-distribution uniformity, these volumetric approaches are rapidly maturing into versatile platforms for generating functional living tissues.
Acoustic-based VBP
Acoustic-based VBP uses focused ultrasound to trigger material solidification deep within a volume, exploiting the superior penetration of sound in opaque media, including tissue. The governing mechanisms typically involve acoustic cavitation or localized sono-thermal heating within a “sono-responsive” bioink, so that polymerization is confined to the acoustic focus129,130. A pioneering realization is direct sound printing (DSP), introduced by Habibi et al.129, wherein a focused transducer generates a microvolume of extreme pressure-temperature via transient cavitation; the collapse of microbubbles initiates polymerization within that focal voxel, and solidifying the focus features in three dimensions. Early DSP established two critical capabilities: curing materials recalcitrant to light- or heat-based methods (including certain thermosets) and remote distance printing (RDP), i.e., writing through intervening opaque barriers. As a single-spot method, however, basic DSP is throughput-limited, analogous to point-scanning optics.
Holographic direct sound printing (HDSP) addresses this limitation by shaping the acoustic field into many foci simultaneously. Derayatifar et al.130 used an acoustic hologram, a designed mask/metamaterial that modulates an incident wavefront, to synthesize a predetermined three-dimensional pressure distribution within the resin. In their system, a stationary hologram generated the patterned field while a robotic stage moved the vat in synchrony so that successive “slices” formed continuously beneath the holographic focus. HDSP retained DSP’s unique advantages and extended them to remotely project patterns through macroscopic slabs of porcine tissue (skin/fat/muscle) to polymerize a helical structure in resin behind the tissue, highlighting a path toward non-invasive in situ printing in vivo.
Without using robotic stage to grow the structure, Kuang et al.131 reported deep-penetration acoustic volumetric printing (DAVP) based on sono-thermal polymerization using a “self-enhancing sono-ink” to yield suspended 3D objects (Fig. 4D). Rather than relying on cavitation chemistry, their bioink combined an ultrasound absorber (to convert acoustic energy to heat), a temperature-sensitive crosslinker (salt trigger), a polymer matrix, and viscosity modifiers for stability. Focused ultrasound scanning created localized hot spots that solidified only at the intersect with the beam focus, enabling three-dimensional writing at centimeter depths and through intervening tissues. Demonstrations included multi-layer honeycombs, branched vascular-like networks, and intricate forms (e.g., a small hand, a spider) printed several millimeters to ~centimeters deep with feature sizes on the order of a few hundred micrometers. Critically, through-tissue printing was validated by placing animal tissue slabs, e.g., ∼17 mm pig liver or composite skin-fat-muscle, above the resin and polymerizing complex shapes beneath. In an ex-vivo cardiac model, sono-ink was injected into a chamber of a goat heart and externally applied ultrasound was used to solidify the material, sealing the left atrial appendage without an incision. A dual-beam configuration further improved precision and safety: two intersecting beams cured only at their overlap, reducing per-beam power, sharpening the effective voxel, and mitigating collateral heating.
In conclusion, acoustic VBP offers capabilities orthogonal to light-based methods: deep, non-contact volumetric curing across opaque barriers and within enclosed or biological environments, rapid parallelization via acoustic holography, and operation with inks that respond to heat or cavitation rather than photochemistry. This strategy remains challenges, tightening the acoustic focal volume (typically ≥ 100 µm), managing acoustic streaming while ensuring uniform cell encapsulation, and expanding robust, cytocompatible sono-responsive inks, are active areas of development132.
Hybrid volumetric bioprinting
Hybrid strategies combine complementary physical mechanisms to achieve continuous, high-throughput biofabrication with conventional materials. Dynamic interface printing (DIP), reported by Vidler et al.133, integrates acoustic modulation with projection photopolymerization at a constrained air-liquid boundary that serves as a deformable build surface (Fig. 4D). A hollow print head traps a thin air layer beneath a transparent window; projected images cure the photocurable bioink at this interface while applied acoustic vibrations induce capillary waves that continually refresh the resin, enhance oxygen/monomer transport, and prevent particle or cell sedimentation. The result is rapid, essentially continuous printing, centimeter-scale structures (e.g., a 5 cm heart) in under 40 s, without exotic photochemistry (standard acrylates and GelMA suffice) or complex optics133. Because the interface is continuously renewed and gently mixed, cell-laden constructs exhibit high viability and homogeneous encapsulation after builds that conclude in seconds. DIP also enables in situ printing by bringing the head to the target surface, straightforward parallelization using multiple heads, improved conversion and mechanical properties at the curing front, and even three-dimensional patterning of suspended particulates by tuning the acoustic frequency to organize inclusions during cure. In this sense, DIP occupies a hybrid niche, neither strictly volumetric nor conventionally layered, combining volumetric-class speed and geometric freedom with simple projection hardware and broadly compatible bioinks.
Together, acoustic and hybrid VBP approaches complement tomographic and multi-beam optical methods by extending volumetric printing into opaque, clinically relevant settings and by enabling continuous, high-throughput fabrication with minimal constraints on ink rheology or optical transparency. As materials, acoustic field-shaping, and thermal management continue to improve, and as hybrid schemes integrate ultrasound’s mixing and transport advantages with light’s spatial precision, these platforms are poised to advance in situ biofabrication and accelerate the translation of volumetric printing toward functional, clinically meaningful tissues.
Challenges and discussion
Concerns on the regulatory of 3D bioprinting assisted NAMs
The regulatory acceptance of 3D bioprinted NAMs is contingent upon addressing several concerns that impact their safety, efficacy, and reliability. These concerns are multifaceted, involving the standardization of materials, the biological fidelity of the models, and the legal frameworks that govern their use. According to the summary of current bioprinted in vitro models from Tables 2–4, the first and perhaps most vital concern is the definition of the COU for a specific 3D printed model. Regulators require a clear statement describing how the NAMs will be utilized, such as for early-stage safety screening, hazard identification, or supporting a point of departure for quantitative risk assessment. Because U.S. federal agencies operate under different statutes, a bioprinted model validated for an EPA pesticide assessment might not be considered acceptable for an FDA drug filing without additional context-specific validation5,29,30.
Another significant concern regarding the biological relevance of 3D printed models, specifically, how closely a test output reflects an effect in the target species, usually humans. Mechanistic understanding is prioritized over simple empirical correlation, and anchoring a bioprinted model to an established AOP provides the biological plausibility needed for regulatory confidence. The difference between 2D and 3D toxicological responses highlights the importance of functional recapitulation5. For example, 3D heart cantilever models have demonstrated sensitivity to titanium dioxide that was entirely missed in 2D cell cultures, as the 3D model was able to reflect the impact of the toxin on spontaneous beating and contractile force134.
Technical characterization is a key component of the regulatory oversight of 3D printed NAMs. This involves assessing every expected source of variability, including the environmental conditions within the printer, the stability of the bioink over time, and the precision of the hardware45. First, the selection and characterization of bioinks are of primary concern in the validation of 3D printed models. Bioinks should function as a nurturing 3D environment that maintains cell phenotype, differentiation potency, and vitality. Technical requirements for bioink rheological and mechanical properties vary by printing method48,80,93. Validation should demonstrate that the bioink can smoothly extrude and rapidly recover its internal structure to retain shape fidelity. Models must be stabilized through crosslinking (e.g., photo-crosslinking or thermal gelation), and regulations require evidence that the crosslinking process does not introduce chemical or physical toxicity to the encapsulated cells. Additionally, natural materials like collagen and alginate71 are prone to variability. Validation protocols should include standardized testing of raw material purity and performance metrics to ensure reproducibility across different study cohorts. The resolution of the printing system determines the model’s ability to replicate fine physiological structures, such as capillary networks or the human lung alveoli. To validate that a 3D printed model is faithful to native tissue, the process should achieve precise deposition and localization of cells in user-defined geometries. This often requires post-print assessment using high-content imaging or histological analysis to confirm the spatial arrangement matches the digital design135. Specific validations for 3D printed models should move beyond cell viability to include tissue-level functional benchmarks. For example, validating vascular constructs requires measuring hemocompatibility136, including platelet adhesion and activation on the printed surfaces136.
The challenge of engineering biomimetic, multi-scale vasculature
While vasculature is not a universal requirement for all in vitro applications, creating a life-sustaining vascular network remains arguably the greatest obstacle to engineering large-scale, functional tissues. Overcoming the diffusion limit (~150–200 µm) is essential to prevent necrotic cores, recapitulate native microenvironments, and enable the study of drug absorption, distribution, metabolism, and excretion relying on endothelial transport18,27. The human circulatory system is a masterpiece of hierarchical design, with vessel diameters spanning multiple orders of magnitude, from centimeter-scale arteries and veins down to 10–15 µm capillaries where essential nutrient and gas exchange occurs137. This multi-scale architecture poses a fundamental manufacturing dilemma for 3D bioprinting. For instance, extrusion-based 3D printing, a dominant strategy in biofabrication, must use wide-bore nozzles to fabricate large-diameter vascular conduits, a choice that inherently sacrifices the resolution needed to print fine capillary beds. Conversely, employing high-resolution nozzles capable of producing micrometer-scale vessels makes the layer-by-layer fabrication of an organ-scale construct prohibitively time-consuming and computationally intensive. This dilemma creates a critical bottleneck that pits fabrication speed against resolution, a trade-off that challenges many additive manufacturing techniques.
Recent progress has begun to address this challenge from two directions: algorithmic design and novel printing modalities. On the design front, powerful new computational platforms have been developed for the rapid, model-guided generation of organ-scale synthetic vasculature. These algorithms, often leveraging principles of fluid dynamics and space-filling optimization, can generate complex, biomimetic vascular tree models for arbitrarily shaped volumes in a matter of minutes, a process that is over 200 times faster than previous methods138. These generated networks can contain millions of vessel segments, ensuring that any cell within the modeled tissue is within ~100–150 µm of a potential blood supply, and the designs can be directly translated into printable instructions138. On the fabrication front, a novel technique has emerged where the extrusion speed of a bioprinter is dynamically varied during printing. This allows a single extruded filament to have a variable diameter along its length, effectively mimicking a tapering blood vessel132.
While these advances are significant, they do not fully resolve the scale-versus-resolution trade-off. The variable-speed technique is demonstrated for a single filament, and its application to a complex, interconnected, and branching network remains a major challenge (Fig. 5). The algorithmic designs are computationally impressive, but their efficient and faithful physical fabrication is the true bottleneck. The solution lies not in a single piece of hardware, but in a more intelligent software and data-integration strategy. The path forward is to combine these advanced vascular generation algorithms with the predictive and organizational power of artificial intelligence (AI), inspiring by recent achievements of utilizing AI models to control the 3D printing quality139,140,141.
The role of AI in this context is transformative. First, AI, particularly through machine learning (ML) and deep learning (DL) approaches, can perform smart path generation. Given a complex vascular architecture, an ML model can analyze the geometry and devise an optimal, time-saving, and non-interfering print path for the bioprinter nozzle, maximizing fabrication efficiency and structural integrity142. Second, and more profoundly, AI can enable hybrid slicing and printing modalities. An intelligent control system could analyze the vascular tree and automatically partition it, designating large-diameter vessels to be printed using a rapid, layer-by-layer slicing method, while seamlessly switching to a high-resolution, trajectory-based method (like variable-speed extrusion) for the delicate, branching capillary networks. This AI-orchestrated hybrid approach would dynamically deploy the most appropriate fabrication technique for each part of the structure, effectively resolving the central conflict between scale and resolution. This reframes the vascularization challenge from a purely mechanical problem to a complex optimization problem perfectly suited for AI-driven solutions.
The intricacy of fabricating heterogeneous anatomical architectures
Beyond vascularization, the second grand challenge lies in constructing the bulk tissue itself, a large-scale, anatomically correct structure composed of multiple, spatially organized cell types and extracellular matrix components (Fig. 5). The leading technology for this task is embedded ink writing (EIW), also known as freeform reversible embedding of suspended hydrogels (FRESH) or embedded bioprinting83. In this process, a functional bioink is extruded from a moving nozzle into a support bath composed of a yield-stress fluid. This bath behaves like a solid matrix, physically holding the delicate printed structure in place, yet it locally and temporarily fluidizes around the nozzle tip under shear stress, permitting true omnidirectional printing of complex, overhanging structures83.
EIW has been instrumental in creating tissues with integrated vascular channels, often by printing a fugitive or sacrificial ink (e.g., gelatin) into a living tissue matrix. This sacrificial network is later evacuated to leave behind a perfusable channel system90,143. This approach has enabled the fabrication of impressive constructs, including anatomic-sized human kidney models and even full-scale human heart models, demonstrating its potential for creating multiscale architectures144,145. However, this strategy is not without significant drawbacks, which become more pronounced as constructs are scaled up to organ size. A primary concern is the “deformation and redundancy” problem. When the vascular network is printed into a pre-formed bulk structure, the process of carving out space for the channels can physically displace and deform the delicate anatomical architecture. Furthermore, the nature of EIW as a serial process, drawing one line at a time, makes it highly inefficient for fabricating large, relatively solid volumes. The printing time increases dramatically with construct size, and the dense, repetitive infill paths required for the bulk tissue can lead to contour deformation and compromise structural fidelity.
This inefficiency reveals two fundamentally different tasks of EIW: the rapid creation of large volumes and the precise placement of intricate features. An emerging EIW-based strategy, “reverse contour printing”82,146, seeks to reconcile these competing requirements by first fabricating the negative space (sacrificial boundary) of the target geometry and then encapsulating it with a bulk material that simultaneously serves as the support bath. After gelation or crosslinking of the encapsulated phase, the sacrificial boundary is removed to reveal the final part. Advancing this approach hinges on two challenges: (i) reliable construction of complex, non-planar, closed-surface negative molds, and (ii) process optimization for rapid, uniform crosslinking throughout the encapsulated bulk material. Furthermore, multi-technology manufacturing illustrates a potential solution for solving the aforementioned challenges by combining volumetric additive manufacturing (VAM) and EIW to fabricate products that are heterogeneous in function, material, and spatial cellular arrangement to better simulate the function and microenvironment for replacing animal models. For instance, embedded extrusion-volumetric printing (EmVP) is a novel hybrid biofabrication technique (CAL + EIW) that merges extrusion bioprinting with ultra-fast, layer-less volumetric printing to shape complex, multicellular, and multi-material tissues147. The process utilizes specially developed light-responsive microgels (µResins) as a support bath, allowing for the precise extrusion of different cell types or materials into the bath. Subsequently, the entire microgel bath is sculpted by tomographic light projections in seconds to form a centimeter-scale, microporous construct. This method enables the creation of regenerative grafts and advanced disease models. The resulting microporous environment has been shown to enhance the differentiation of various stem cells (vascular, mesenchymal, and neural). As a proof-of-concept, EmVP was used to create a synthetic biology model where adipocyte differentiation was controlled by optogenetically engineered pancreatic cells, showcasing its potential for developing complex, functional living systems. However, this hybrid system is fundamentally constrained by an inherent limitation. Its typical operation involves batch-wise rotation, a method that inherently hinders the establishment of a truly continuous manufacturing workflow, thereby limiting its efficiency in very high-volume production scenarios.
Therefore, an interesting perspective is to synergistically combine the aforementioned VAM technologies and advanced EIW methods to realize a continuous flow of fabrication and fully automated operation. This would combine the strengths of two systems: VAM, excelling at the near-instantaneous fabrication of entire solid objects, and EIW, which is ideal for the precise, multi-material deposition of functional cell groups and high-resolution vascular networks. Simultaneously, this hybrid system will call for the development of innovative materials that possess the supporting function for EIW, the photocurable property required for VAM, and an appropriate logic design (such as “AND” or “AND NOT” gates126) to support the continuous flow of the combined systems. More active components will also be needed to enhance light penetration, improve printing resolution, and increase biocompatibility. Overall, hybrid systems will offer a broad selection of materials and enable more complicated printed geometries and intricate structures, generating highly biomimetic organoids to serve as alternative animal models for drug testing and disease research.
The challenge of functionalizing in vitro models with biochemical cues, cellular diversity, and multi-organ interactions
While the faithful replication of the mechanical environment represents one challenge in tissue engineering, a parallel and critical frontier lies in endowing in vitro models with the rich biochemical complexity, cellular diversity, and systemic interconnectedness of living organisms. Tissues do not exist in isolation; they are complex ecosystems defined by steep chemical gradients, a dynamic interplay between diverse cell populations, and constant communication with distant organs. To advance beyond simple organ-level mimicry toward true physiological relevance, particularly at host-microbiome interfaces, engineered systems must capture these multifaceted biological realities (Fig. 5). This chapter outlines the key challenges and innovative engineering solutions aimed at integrating the essential biochemical cues, cellular components, and multi-organ interactions that govern tissue function, homeostasis, and disease.
An obstacle in modeling densely colonized tissues is recreating the different environmental requirements of aerobic host cells and anaerobic or microaerophilic microbes148. The steep physicochemical gradients, particularly in oxygen and pH, that define these niches are critical for shaping microbial composition and host responses, yet they are notoriously difficult to reproduce in vitro148. Engineering solutions to establish and maintain these steep oxygen gradients vary in complexity. Sealed transwell systems offer a straightforward approach by physically separating microbial and host components with a porous membrane, maintaining anoxia in the apical compartment while oxygenated media support the host cells basally149. This strategy has been used to support co-cultures of colonic organoid monolayers with anaerobic commensals. For more complex gradient control, biomaterial-based systems, such as porous silk150 or gelatin methacryloyl scaffolds151, can achieve oxygen depletion through the autonomous consumption by embedded host epithelial cells. Microfluidic devices152 can provide the dynamic control over oxygen gradients while also addressing nutrient depletion through continuous perfusion. These models often feature parallel microchannels separated by a thin membrane, allowing oxygenated media to be perfused through the host cell compartment while deoxygenated media flows through the microbial channel, achieving oxygen concentrations below 0.5% in the luminal space152.
Similarly, spatial pH gradients create distinct niches that promote or inhibit the growth of specific microbes. In organ-on-a-chip models of the vagina and cervix, for example, physiologically acidic conditions (pH ~4.0)153 have been established through the controlled perfusion of pH-adjusted media, successfully sustaining colonization by commensal lactobacilli and mirroring clinical observations of dysbiosis. A more sophisticated approach154 for modeling the gut has utilized acid-producing bacteria embedded within a multi-layered hydrogel, where distinct bacterial compositions within each layer metabolize nutrients to generate a stable pH gradient without external control, enabling the study of patient-derived microbiota on drug efficacy.
Native tissues are not monocultures but are composed of a heterogeneous mix of cell types embedded within a complex and dynamic extracellular matrix (ECM). Replicating this multicellularity and the essential structural and biochemical cues provided by the ECM is critical for achieving functional fidelity. The ECM is a central regulator of cell behavior155, and its omission creates a significant fidelity gap, especially as commensal bacteria can directly degrade various host ECM components156. To address this, researchers employ strategies ranging from surface coatings of ECM-derived materials like collagen I to promote cell attachment, to using ECM-based materials as 3D hydrogel substrates that encapsulate cells. For instance, in a gingiva-on-a-chip model157, primary human gingival fibroblasts seeded within a 3D fibrin-based matrix facilitated the formation of a stratified gingival epithelium. Furthermore, to capture the complex interplay between host cell types, advanced models increasingly integrate cellular diversity using primary cells, which preserve native responsiveness, or induced pluripotent stem cells (iPSCs) to generate diverse cell types like neurons158. These sources can be combined in co-culture strategies within 3D hydrogels or across porous membranes. Organoids and tissue explants provide an additional layer of relevance by preserving native tissue architecture and can be integrated into microfluidic chips to make their luminal surfaces accessible.
The influence of the microbiome extends beyond its local environment, with microbial metabolites and immune signals exerting broad systemic effects on distant organs. Single-tissue models cannot capture this systemic complexity, driving the development of multi-organ platforms148. Immune-competent gut models, for example, incorporate various immune cell populations to study how the host response is modulated. The co-culture of colonic epithelial cells with antigen-presenting cells and T cells revealed that the pro-inflammatory response to the commensal Faecalibacterium prausnitzii was attenuated by naïve T cells, demonstrating a mechanism of immune tolerance159. Other work has shown that integrating macrophages and dendritic cells supports stable colonization by probiotics and prevents pathogen overgrowth160. Beyond local interactions, microfluidic platforms are uniquely suited for modeling multi-organ communication by connecting distinct, individually controlled tissue compartments. A system designed to study gut-liver-immune interactions demonstrated that lipopolysaccharide (LPS)-treated macrophages triggered a pro-inflammatory response that propagated to the liver161. Similarly, the gut-brain axis has been modeled using dual-perfusion devices that connect gut and brain compartments, showing that extracellular vesicles derived from commensal bacteria can translocate from the intestine and enter the neural compartment162.
To potentially solve aforementioned challenges, advanced 3D bioprinting offers a transformative platform for engineering in vitro tissue models that more faithfully recapitulate the biochemical, cellular, and systemic complexity of native tissues. By integrating data-driven bioink design163,164 with multi-material printing strategies, bioprinting enables the fabrication of heterogeneous constructs165,166 in which diverse cell populations are precisely organized within tunable extracellular matrices, thereby replicating the spatial architecture and functional integration of in vivo environments. The technique further allows intrinsic engineering of physicochemical gradients, such as oxygen and pH profiles, through controlled construct geometry, porosity, and cell density, and can be seamlessly combined with microfluidics to establish dynamic host-microbe interfaces. Extending beyond single-tissue systems, multi-material bioprinting167,168 supports the construction of interconnected organotypic modules with vascular-like channels, permitting the study of systemic inter-organ communication, such as microbiome-derived metabolite transport and immune-mediated signaling. Therefore, these advances position bioprinting as a critical enabler of physiologically relevant, multi-scale models for studying health and disease mechanisms.
Conclusion and outlook
The FDA Modernization Act 2.0 has accelerated demand for validated NAMs that anticipate human responses more reliably than traditional animal models. Within this landscape, 3D bioprinting now represents a premier class of tools for fabricating systematic, human-derived tissue models. Established methods, especially for inkjet bioprinting and extrusion-based printing, are already enabling layered epithelia (e.g., skin, airway) and larger constructs with perfusable channels, balancing resolution, throughput, and material compatibility. Vat photopolymerization-based approaches, including two-photon polymerization and digital light processing, add micro-architectural precision and speed, supporting organotypic units such as hepatic lobules and tumor models that capture essential biomechanical and cellular diversity.
Volumetric bioprinting defines the current frontier. Light-driven methods such as Computed axial lithography and Xolography can form complex, cell-laden geometries within seconds while preserving viability. Acoustic volumetric strategies further expand the toolbox by operating through optically opaque media, raising the prospect of non-invasive, in situ fabrication. Hybrid systems combine complementary physical mechanisms to push both speed and versatility.
Key barriers remain before organ-scale function becomes routine. Four challenges dominate: (i) calling for specific regulations for 3D bioprinted in vitro models, (ii) engineering hierarchical, biomimetic vasculature that sustains thick tissues, (iii) building anatomically faithful, heterogeneous architectures across scales, and (iv) functionalizing in vitro models with biochemical cues, cellular diversity, and multi-organ interactions to create models of unprecedented fidelity. Progress will hinge on coupling advanced manufacturing with intelligent design, using AI-guided process and material optimization, and platforms that dynamically switch between high-speed and high-resolution modes to reconcile scale with detail. Strategic combinations, for example, employing volumetric additive methods for rapid bulk shaping followed by fine-feature and multi-material techniques (e.g., embedded ink writing) for microstructure, are poised to yield organoids and tissue models with unprecedented fidelity. Continued innovation along these lines aligns squarely with the intent of the FDA modernization effort: a more ethical, efficient, and critically predictive preclinical ecosystem that shortens timelines, reduces attrition, and delivers safer, more effective therapies to patients.
Data availability
No datasets were generated or analyzed during the current study.
References
-
Hajar, R. Animal testing and medicine. Heart Views 12, 42, https://doi.org/10.4103/1995-705X.81548 (2011).
-
Sun, D., Gao, W., Hu, H. & Zhou, S. Why 90% of clinical drug development fails and how to improve it?. Acta Pharm. Sin. B 12, 3049–3062, https://doi.org/10.1016/j.apsb.2022.02.002 (2022).
-
Hay, M., Thomas, D. W., Craighead, J. L., Economides, C. & Rosenthal, J. Clinical development success rates for investigational drugs. Nat. Biotechnol. 32, 40–51, https://doi.org/10.1038/nbt.2786 (2014).
-
Arrowsmith, J. & Miller, P. Trial watch: phase II and phase III attrition rates 2011-2012. Nat. Rev. Drug Discov. 12, 569, https://doi.org/10.1038/nrd4090 (2013).
-
Ouedraogo, G., Alepee, N., Tan, B. & Roper, C. S. A call to action: Advancing new approach methodologies (NAMs) in regulatory toxicology through a unified framework for validation and acceptance. Regul. Toxicol. Pharm. 162, 105904, https://doi.org/10.1016/j.yrtph.2025.105904 (2025).
-
Marx, U. et al. Biology-inspired microphysiological systems to advance patient benefit and animal welfare in drug development. ALTEX 37, 365–394, https://doi.org/10.14573/altex.2001241 (2020).
-
Low, L. A., Mummery, C., Berridge, B. R., Austin, C. P. & Tagle, D. A. Organs-on-chips: into the next decade. Nat. Rev. Drug Discov. 20, 345–361, https://doi.org/10.1038/s41573-020-0079-3 (2021).
-
Congress, U. S. & H.R. Consolidated Appropriations Act, 2023 (including FDA Modernization Act 2.0. Pub L, 117–328 (2022).
-
Wadman, M. FDA no longer has to require animal testing for new drugs. Science 379, 127–128, https://doi.org/10.1126/science.adg6276 (2023).
-
U.S. Food & Drug Administration. Roadmap to Reducing Animal Testing in Preclinical Safety Studies. (U.S. Food & Drug Administration, 2025).
-
Ma, L. D. et al. Design and fabrication of a liver-on-a-chip platform for convenient, highly efficient, and safe in situ perfusion culture of 3D hepatic spheroids. Lab Chip 18, 2547–2562, https://doi.org/10.1039/c8lc00333e (2018).
-
Toepke, M. W. & Beebe, D. J. PDMS absorption of small molecules and consequences in microfluidic applications. Lab Chip 6, 1484–1486, https://doi.org/10.1039/b612140c (2006).
-
Ingber, D. E. Human organs-on-chips for disease modelling, drug development and personalized medicine. Nat. Rev. Genet. 23, 467–491, https://doi.org/10.1038/s41576-022-00466-9 (2022).
-
Skardal, A., Shupe, T. & Atala, A. Organoid-on-a-chip and body-on-a-chip systems for drug screening and disease modeling. Drug Discov. Today 21, 1399–1411, https://doi.org/10.1016/j.drudis.2016.07.003 (2016).
-
Ronaldson-Bouchard, K. & Vunjak-Novakovic, G. Organs-on-a-chip: a fast track for engineered human tissues in drug development. Cell Stem Cell 22, 310–324, https://doi.org/10.1016/j.stem.2018.02.011 (2018).
-
Mandrycky, C., Phong, K. & Zheng, Y. Tissue engineering toward organ-specific regeneration and disease modeling. MRS Commun. 7, 332–347, https://doi.org/10.1557/mrc.2017.58 (2017).
-
Schmeichel, K. L. & Bissell, M. J. Modeling tissue-specific signaling and organ function in three dimensions. J. Cell Sci. 116, 2377–2388, https://doi.org/10.1242/jcs.00503 (2003).
-
Jain, R. K., Au, P., Tam, J., Duda, D. G. & Fukumura, D. Engineering vascularized tissue. Nat. Biotechnol. 23, 821–823, https://doi.org/10.1038/nbt0705-821 (2005).
-
Ren, Y. et al. Developments and opportunities for 3D bioprinted organoids. Int J. Bioprint 7, 364, https://doi.org/10.18063/ijb.v7i3.364 (2021).
-
Banchini, F. et al. 3-D reconstruction in liver surgery: a systematic review. HPB 26, 1205–1215, https://doi.org/10.1016/j.hpb.2024.06.006 (2024).
-
Clark, A. M., Allbritton, N. L. & Wells, A. Integrative microphysiological tissue systems of cancer metastasis to the liver. Semin Cancer Biol. 71, 157–169, https://doi.org/10.1016/j.semcancer.2020.06.010 (2021).
-
da Silva, V. A. et al. Building the framework for bioprinted human heart tissue: recent developments and future prospects. Regen. Med. 20, 409–430, https://doi.org/10.1080/17460751.2025.2558269 (2025).
-
Ronaldson-Bouchard, K. et al. A multi-organ chip with matured tissue niches linked by vascular flow. Nat. Biomed. Eng. 6, 351–371, https://doi.org/10.1038/s41551-022-00882-6 (2022).
-
Avula, L. R. & Grodzinski, P. How organ-on-a-chip is advancing cancer research and oncology – a cancer hallmarks’ perspective. Front. Lab Chip Technol. 3, https://doi.org/10.3389/frlct.2024.1487377 (2024).
-
Mandrycky, C., Wang, Z., Kim, K. & Kim, D. H. 3D bioprinting for engineering complex tissues. Biotechnol. Adv. 34, 422–434, https://doi.org/10.1016/j.biotechadv.2015.12.011 (2016).
-
Donderwinkel, I., van Hest, J. C. M. & Cameron, N. R. Bio-inks for 3D bioprinting: recent advances and future prospects. Polym. Chem. 8, 4451–4471, https://doi.org/10.1039/c7py00826k (2017).
-
Gold, K., Gaharwar, A. K. & Jain, A. Emerging trends in multiscale modeling of vascular pathophysiology: Organ-on-a-chip and 3D printing. Biomaterials 196, 2–17, https://doi.org/10.1016/j.biomaterials.2018.07.029 (2019).
-
Murphy, S. V. & Atala, A. 3D bioprinting of tissues and organs. Nat. Biotechnol. 32, 773–785, https://doi.org/10.1038/nbt.2958 (2014).
-
Mirlohi, M. S., Yousefi, T., Aref, A. R. & Seyfoori, A. Integrating new approach methodologies (NAMs) into preclinical regulatory evaluation of oncology drugs. Biomimetics 10, https://doi.org/10.3390/biomimetics10120796 (2025).
-
Sewell, F. et al. New approach methodologies (NAMs): identifying and overcoming hurdles to accelerated adoption. Toxicol. Res. 13, tfae044. https://doi.org/10.1093/toxres/tfae044 (2024).
-
Albritton, J. L. & Miller, J. S. 3D bioprinting: improving in vitro models of metastasis with heterogeneous tumor microenvironments. Dis. Model Mech. 10, 3–14, https://doi.org/10.1242/dmm.025049 (2017).
-
Yi, H. G. et al. Application of 3D bioprinting in the prevention and the therapy for human diseases. Signal Transduct. Target Ther. 6, 177, https://doi.org/10.1038/s41392-021-00566-8 (2021).
-
Teli, P., Kale, V. & Vaidya, A. Beyond animal models: revolutionizing neurodegenerative disease modeling using 3D in vitro organoids, microfluidic chips, and bioprinting. Cell Tissue Res 394, 75–91, https://doi.org/10.1007/s00441-023-03821-2 (2023).
-
Gungor-Ozkerim, P. S., Inci, I., Zhang, Y. S., Khademhosseini, A. & Dokmeci, M. R. Bioinks for 3D bioprinting: an overview. Biomater. Sci. 6, 915–946, https://doi.org/10.1039/c7bm00765e (2018).
-
Deo, K. A., Singh, K. A., Peak, C. W., Alge, D. L. & Gaharwar, A. K. Bioprinting 101: design, fabrication, and evaluation of cell-laden 3D bioprinted scaffolds. Tissue Eng. Part A 26, 318–338, https://doi.org/10.1089/ten.TEA.2019.0298 (2020).
-
Lipsky, M. S. & Sharp, L. K. From idea to market: the drug approval process. J. Am. Board Fam. Pr. 14, 362–367 (2001).
-
Hughes, J. P., Rees, S., Kalindjian, S. B. & Philpott, K. L. Principles of early drug discovery. Br. J. Pharm. 162, 1239–1249, https://doi.org/10.1111/j.1476-5381.2010.01127.x (2011).
-
Kang, D. et al. All-inkjet-printed 3D alveolar barrier model with physiologically relevant microarchitecture. Adv. Sci. 8, 2004990. https://doi.org/10.1002/advs.202004990 (2021).
-
Lee, H.-R. et al. 3D microextrusion-inkjet hybrid printing of structured human skin equivalents. Bioprinting 22, 00143, https://doi.org/10.1016/j.bprint.2021.e00143 (2021).
-
Wu, D. W. et al. Bioprinting of perfusable vascularized organ models for drug development via sacrificial-free direct ink writing. Adv. Funct. Mater. 34, 2314171. https://doi.org/10.1002/adfm.202314171 (2024).
-
Hwang, K. S. et al. Augmented 3D Printing for Multiscale Microphysiological Systems. Small, e2504750, https://doi.org/10.1002/smll.202504750 (2025).
-
Rexius-Hall, M. L., Madrigal, M. D., Kilic, C. Y., Shen, K. & McCain, M. L. Profiling paracrine interactions between hypoxic and normoxic skeletal muscle tissue in a microphysiological system fabricated from 3D printed components. Lab Chip 25, 212–224, https://doi.org/10.1039/d4lc00603h (2025).
-
Schreiber, K. et al. Challenges of designing and conducting cohort studies and clinical trials in populations of pregnant people. Lancet Rheumatol. 6, e560–e572, https://doi.org/10.1016/S2665-9913(24)00118-8 (2024).
-
Tome-Carneiro, J. et al. Resveratrol and clinical trials: the crossroad from in vitro studies to human evidence. Curr. Pharm. Des. 19, 6064–6093, https://doi.org/10.2174/13816128113199990407 (2013).
-
Iccvam. Validation, Qualification, and Regulatory Acceptance of New Approach Methodologies. (2024). https://doi.org/10.22427/niceatm-2
-
Xie, R. et al. A comprehensive review on 3D tissue models: biofabrication technologies and preclinical applications. Biomaterials 304, 122408. https://doi.org/10.1016/j.biomaterials.2023.122408 (2024).
-
Park, J. A., Lee, Y. & Jung, S. Inkjet-based bioprinting for tissue engineering. Organoid 3, 12, https://doi.org/10.51335/organoid.2023.3.e12 (2023).
-
Zhang, Z. et al. Evaluation of bioink printability for bioprinting applications. Appl. Phys. Rev. 5, https://doi.org/10.1063/1.5053979 (2018).
-
Saunders, R. E. & Derby, B. Inkjet printing biomaterials for tissue engineering: bioprinting. Int. Mater. Rev. 59, 430–448, https://doi.org/10.1179/1743280414y.0000000040 (2014).
-
Shi, J. et al. Shear stress analysis and its effects on cell viability and cell proliferation in drop-on-demand bioprinting. Biomed. Phys. Eng. Express 4, https://doi.org/10.1088/2057-1976/aac946 (2018).
-
Ng, W. L. & Shkolnikov, V. Jetting-based bioprinting: process, dispense physics, and applications. Bio-Des. Manuf. 7, 771–799, https://doi.org/10.1007/s42242-024-00285-3 (2024).
-
Lim, K. S. et al. Fundamentals and applications of photo-cross-linking in bioprinting. Chem. Rev. 120, 10662–10694, https://doi.org/10.1021/acs.chemrev.9b00812 (2020).
-
Nie, L. et al. Click chemistry for 3D bioprinting. Mater. Horiz. 10, 2727–2763, https://doi.org/10.1039/d3mh00516j (2023).
-
Parak, A. et al. Functionalizing bioinks for 3D bioprinting applications. Drug Discov. Today 24, 198–205, https://doi.org/10.1016/j.drudis.2018.09.012 (2019).
-
Garciamendez-Mijares, C. E. et al. Acoustic bioprinting: a glimpse into an emerging field. Small Methods, e2500733, https://doi.org/10.1002/smtd.202500733 (2025).
-
Naranjo-Alcazar, R., Bendix, S., Groth, T. & Gallego Ferrer, G. Research progress in enzymatically cross-linked hydrogels as injectable systems for bioprinting and tissue engineering. Gels 9, https://doi.org/10.3390/gels9030230 (2023).
-
GhavamiNejad, A., Ashammakhi, N., Wu, X. Y. & Khademhosseini, A. Crosslinking strategies for 3D bioprinting of polymeric hydrogels. Small 16, e2002931, https://doi.org/10.1002/smll.202002931 (2020).
-
Mironi-Harpaz, I., Wang, D. Y., Venkatraman, S. & Seliktar, D. Photopolymerization of cell-encapsulating hydrogels: crosslinking efficiency versus cytotoxicity. Acta Biomater. 8, 1838–1848, https://doi.org/10.1016/j.actbio.2011.12.034 (2012).
-
Rozbesky, D. et al. Impact of chemical cross-linking on protein structure and function. Anal. Chem. 90, 1104–1113, https://doi.org/10.1021/acs.analchem.7b02863 (2018).
-
MJ, L. D. et al. The impact of the ionic cross-linking mode on the physical and in vitro dexamethasone release properties of chitosan/hydroxyapatite beads. Molecules 24, https://doi.org/10.3390/molecules24244510 (2019).
-
Sardari, M., Ghanati, F., Mobasheri, H. & Hajnorouzi, A. Sound waves alter the viability of tobacco cells via changes in cytosolic calcium, membrane integrity, and cell wall composition. PLoS One 19, e0299055, https://doi.org/10.1371/journal.pone.0299055 (2024).
-
Mazzoccoli, J. P., Feke, D. L., Baskaran, H. & Pintauro, P. N. Development of multilayered cell-hydrogel composites using an acoustic focusing technique. Biotechnol. Prog. 26, 600–605, https://doi.org/10.1002/btpr.332 (2010).
-
Rosendahl, J., Zarna, C., Hakansson, J. & Chinga-Carrasco, G. Gene-expression analysis of human fibroblasts affected by 3D-printed carboxylated nanocellulose constructs. Bioengineering 10, https://doi.org/10.3390/bioengineering10010121 (2023).
-
Li, X. et al. Inkjet bioprinting of biomaterials. Chem. Rev. 120, 10793–10833, https://doi.org/10.1021/acs.chemrev.0c00008 (2020).
-
Kim, W. et al. 3D inkjet-bioprinted lung-on-a-chip. ACS Biomater. Sci. Eng. 9, 2806–2815, https://doi.org/10.1021/acsbiomaterials.3c00089 (2023).
-
Yoon, W. H. et al. Use of inkjet-printed single cells to quantify intratumoral heterogeneity. Biofabrication 12, 035030, https://doi.org/10.1088/1758-5090/ab9491 (2020).
-
Kang, D., Lee, Y., Kim, W., Lee, H. R. & Jung, S. 3D pulmonary fibrosis model for anti-fibrotic drug discovery by inkjet-bioprinting. Biomed. Mater. 18, https://doi.org/10.1088/1748-605X/aca8e3 (2022).
-
Saadi, M. et al. Direct ink writing: a 3D printing technology for diverse materials. Adv. Mater. 34, e2108855. https://doi.org/10.1002/adma.202108855 (2022).
-
Lewis, J. A. Direct ink writing of 3D functional materials. Adv. Funct. Mater. 16, 2193–2204, https://doi.org/10.1002/adfm.200600434 (2006).
-
Ouyang, L., Yao, R., Zhao, Y. & Sun, W. Effect of bioink properties on printability and cell viability for 3D bioplotting of embryonic stem cells. Biofabrication 8, 035020, https://doi.org/10.1088/1758-5090/8/3/035020 (2016).
-
Chen, X. B. et al. Biomaterials / bioinks and extrusion bioprinting. Bioact. Mater. 28, 511–536, https://doi.org/10.1016/j.bioactmat.2023.06.006 (2023).
-
Shakur, M. S. et al. Effect of hydrodynamic shear stress on algal cell fate in 3D extrusion bioprinting. Adv. Eng. Mater. 27, https://doi.org/10.1002/adem.202401768 (2025).
-
Zhang, C. et al. Sacrificial scaffold-assisted direct ink writing of engineered aortic valve prostheses. Biofabrication 15, 045015, https://doi.org/10.1088/1758-5090/aceffb (2023).
-
Tay, R. Y., Song, Y., Yao, D. R. & Gao, W. Direct-ink-writing 3D-printed bioelectronics. Mater. Today 71, 135–151, https://doi.org/10.1016/j.mattod.2023.09.006 (2023).
-
Garcia-Lamas, L. et al. Enriched mesoporous bioactive glass scaffolds as bone substitutes in critical diaphyseal bone defects in rabbits. Acta Biomater. 180, 104–114, https://doi.org/10.1016/j.actbio.2024.04.005 (2024).
-
Zandrini, T., Florczak, S., Levato, R. & Ovsianikov, A. Breaking the resolution limits of 3D bioprinting: future opportunities and present challenges. Trends Biotechnol. 41, 604–614, https://doi.org/10.1016/j.tibtech.2022.10.009 (2023).
-
Wu, Q. et al. Embedded extrusion printing in yield-stress-fluid baths. Matter-Us 5, 3775–3806, https://doi.org/10.1016/j.matt.2022.09.003 (2022).
-
Zhou, K., Sun, Y., Yang, J., Mao, H. & Gu, Z. Hydrogels for 3D embedded bioprinting: a focused review on bioinks and support baths. J. Mater. Chem. B 10, 1897–1907, https://doi.org/10.1039/d1tb02554f (2022).
-
Xie, Z. T., Kang, D. H. & Matsusaki, M. Resolution of 3D bioprinting inside bulk gel and granular gel baths. Soft Matter 17, 8769–8785, https://doi.org/10.1039/d1sm00926e (2021).
-
Hua, W. et al. Fluid bath-assisted 3D printing for biomedical applications: from pre- to postprinting stages. ACS Biomater. Sci. Eng. 7, 4736–4756, https://doi.org/10.1021/acsbiomaterials.1c00910 (2021).
-
Compaan, A. M., Song, K. & Huang, Y. Gellan fluid gel as a versatile support bath material for fluid extrusion bioprinting. ACS Appl Mater. Interfaces 11, 5714–5726, https://doi.org/10.1021/acsami.8b13792 (2019).
-
Hua, W. et al. 3D printing-based full-scale human brain for diverse applications. Brain-X 1, https://doi.org/10.1002/brx2.5 (2023).
-
Hinton, T. J. et al. Three-dimensional printing of complex biological structures by freeform reversible embedding of suspended hydrogels. Sci. Adv. 1, e1500758. https://doi.org/10.1126/sciadv.1500758 (2015).
-
Shiwarski, D. J., Hudson, A. R., Tashman, J. W. & Feinberg, A. W. Emergence of FRESH 3D printing as a platform for advanced tissue biofabrication. APL Bioeng. 5, 010904. https://doi.org/10.1063/5.0032777 (2021).
-
Mirdamadi, E., Tashman, J. W., Shiwarski, D. J., Palchesko, R. N. & Feinberg, A. W. FRESH 3D bioprinting a full-size model of the human heart. ACS Biomater. Sci. Eng. 6, 6453–6459, https://doi.org/10.1021/acsbiomaterials.0c01133 (2020).
-
Bhattacharjee, T. et al. Writing in the granular gel medium. Sci. Adv. 1, e1500655. https://doi.org/10.1126/sciadv.1500655 (2015).
-
Kolesky, D. B. et al. 3D bioprinting of vascularized, heterogeneous cell-laden tissue constructs. Adv. Mater. 26, 3124–3130, https://doi.org/10.1002/adma.201305506 (2014).
-
Patricio, S. G. et al. Freeform 3D printing using a continuous viscoelastic supporting matrix. Biofabrication 12, 035017, https://doi.org/10.1088/1758-5090/ab8bc3 (2020).
-
Noor, N. et al. 3D printing of personalized thick and perfusable cardiac patches and hearts. Adv. Sci. 6, 1900344. https://doi.org/10.1002/advs.201900344 (2019).
-
Skylar-Scott, M. A. et al. Biomanufacturing of organ-specific tissues with high cellular density and embedded vascular channels. Sci. Adv. 5, eaaw2459. https://doi.org/10.1126/sciadv.aaw2459 (2019).
-
Ma, J., Wang, Y. & Liu, J. Bioprinting of 3D tissues/organs combined with microfluidics. RSC Adv. 8, 21712–21727, https://doi.org/10.1039/c8ra03022g (2018).
-
Felipe-Mendes, C., Ruiz-Rubio, L. & Vilas-Vilela, J. L. Biomaterials obtained by photopolymerization: from UV to two photon. Emergent Mater. 3, 453–468, https://doi.org/10.1007/s42247-020-00114-0 (2020).
-
Li, H. et al. Digital light processing (DLP)-based (bio)printing strategies for tissue modeling and regeneration. Aggregate 4, 270, https://doi.org/10.1002/agt2.270 (2023).
-
Alparslan, C. & Bayraktar, S. Advances in digital light processing (DLP) bioprinting: a review of biomaterials and its applications, innovations, challenges, and future perspectives. Polymers 17, https://doi.org/10.3390/polym17091287 (2025).
-
Jing, S. et al. Advances in volumetric bioprinting. Biofabrication 16, 012004, https://doi.org/10.1088/1758-5090/ad0978 (2023).
-
Bernal, P. N. et al. The road ahead in materials and technologies for volumetric 3D printing. Nat. Rev. Mater. 10, 826–841, https://doi.org/10.1038/s41578-025-00785-3 (2025).
-
He, N. et al. Photoinhibiting via simultaneous photoabsorption and free-radical reaction for high-fidelity light-based bioprinting. Nat. Commun. 14, 3063. https://doi.org/10.1038/s41467-023-38838-2 (2023).
-
Huang, Z., Chi-Pong Tsui, G., Deng, Y. & Tang, C.-Y. Two-photon polymerization nanolithography technology for fabrication of stimulus-responsive micro/nano-structures for biomedical applications. Nanotechnol. Rev. 9, 1118–1136, https://doi.org/10.1515/ntrev-2020-0073 (2020).
-
Mao, Q. et al. Fabrication of liver microtissue with liver decellularized extracellular matrix (dECM) bioink by digital light processing (DLP) bioprinting. Mater. Sci. Eng. C. Mater. Biol. Appl 109, 110625. https://doi.org/10.1016/j.msec.2020.110625 (2020).
-
Otuka, A. J. G., Tomazio, N. B., Paula, K. T. & Mendonca, C. R. Two-photon polymerization: functionalized microstructures, micro-resonators, and bio-scaffolds. Polymers 13, https://doi.org/10.3390/polym13121994 (2021).
-
Marino, A., Filippeschi, C., Mattoli, V., Mazzolai, B. & Ciofani, G. Biomimicry at the nanoscale: current research and perspectives of two-photon polymerization. Nanoscale 7, 2841–2850, https://doi.org/10.1039/c4nr06500j (2015).
-
Dobos, A. et al. On-chip high-definition bioprinting of microvascular structures. Biofabrication 13, 015016, https://doi.org/10.1088/1758-5090/abb063 (2021).
-
Dobos, A. et al. Thiol-gelatin-norbornene bioink for laser-based high-definition bioprinting. Adv. Health. Mater. 9, e1900752. https://doi.org/10.1002/adhm.201900752 (2020).
-
Sharaf, A. et al. Two-photon polymerization of 2.5D and 3D microstructures fostering a ramified resting phenotype in primary microglia. Front. Bioeng. Biotechnol. 10, 926642. https://doi.org/10.3389/fbioe.2022.926642 (2022).
-
Mandt, D. et al. Fabrication of biomimetic placental barrier structures within a microfluidic device utilizing two-photon polymerization. Int. J. Bioprint 4, 144, https://doi.org/10.18063/IJB.v4i2.144 (2018).
-
Enrico, A. et al. 3D microvascularized tissue models by laser-based cavitation molding of collagen. Adv. Mater. 34, e2109823. https://doi.org/10.1002/adma.202109823 (2022).
-
Li, W. et al. Stereolithography apparatus and digital light processing-based 3D bioprinting for tissue fabrication. iScience 26, 106039. https://doi.org/10.1016/j.isci.2023.106039 (2023).
-
Goodarzi Hosseinabadi, H., Dogan, E., Miri, A. K. & Ionov, L. Digital light processing bioprinting advances for microtissue models. ACS Biomater. Sci. Eng. 8, 1381–1395, https://doi.org/10.1021/acsbiomaterials.1c01509 (2022).
-
Miri, A. K. et al. Effective bioprinting resolution in tissue model fabrication. Lab Chip 19, 2019–2037, https://doi.org/10.1039/c8lc01037d (2019).
-
Bhusal, A. et al. Multi-material digital light processing bioprinting of hydrogel-based microfluidic chips. Biofabrication 14, https://doi.org/10.1088/1758-5090/ac2d78 (2021).
-
Xu, F. et al. Digital light processing printed hydrogel scaffolds with adjustable modulus. Sci. Rep. 14, 15695. https://doi.org/10.1038/s41598-024-66507-x (2024).
-
Ma, X. et al. Deterministically patterned biomimetic human iPSC-derived hepatic model via rapid 3D bioprinting. Proc. Natl. Acad. Sci. USA 113, 2206–2211, https://doi.org/10.1073/pnas.1524510113 (2016).
-
Zhong, Z. et al. Rapid 3D bioprinting of a multicellular model recapitulating pterygium microenvironment. Biomaterials 282, 121391. https://doi.org/10.1016/j.biomaterials.2022.121391 (2022).
-
Tang, M. et al. Rapid 3D bioprinting of glioblastoma model mimicking native biophysical heterogeneity. Small 17, e2006050. https://doi.org/10.1002/smll.202006050 (2021).
-
Yang, M. et al. Multi-material digital light processing (DLP) bioprinting of heterogeneous hydrogel constructs with perfusable networks. Adv. Funct. Mater. 34, https://doi.org/10.1002/adfm.202316456 (2024).
-
Madrid-Wolff, J. et al. A review of materials used in tomographic volumetric additive manufacturing. MRS Commun. 13, 764–785, https://doi.org/10.1557/s43579-023-00447-x (2023).
-
Whyte, D. J., Doeven, E. H., Sutti, A., Kouzani, A. Z. & Adams, S. D. Volumetric additive manufacturing: a new frontier in layer-less 3D printing. Addit. Manuf. 84, 104094. https://doi.org/10.1016/j.addma.2024.104094 (2024).
-
Mathur, V., Dsouza, V., Srinivasan, V. & Vasanthan, K. S. Volumetric additive manufacturing for cell printing: bridging industry adaptation and regulatory frontiers. ACS Biomater. Sci. Eng. 11, 156–181, https://doi.org/10.1021/acsbiomaterials.4c01837 (2025).
-
Kelly, B. E. et al. Volumetric additive manufacturing via tomographic reconstruction. Science 363, 1075–1079, https://doi.org/10.1126/science.aau7114 (2019).
-
Kim, D., Kang, D., Kim, D. & Jang, J. Volumetric bioprinting strategies for creating large-scale tissues and organs. MRS Bull. 48, 657–667, https://doi.org/10.1557/s43577-023-00541-4 (2023).
-
Toombs, J. T. et al. Volumetric additive manufacturing of silica glass with microscale computed axial lithography. Science 376, 308–312, https://doi.org/10.1126/science.abm6459 (2022).
-
Alvarez-Castano, M. I. et al. Holographic tomographic volumetric additive manufacturing. Nat. Commun. 16, 1551. https://doi.org/10.1038/s41467-025-56852-4 (2025).
-
Rizzo, R., Ruetsche, D., Liu, H. & Zenobi-Wong, M. Optimized Photoclick (Bio)Resins for Fast Volumetric Bioprinting. Adv. Mater. 33, e2102900. https://doi.org/10.1002/adma.202102900 (2021).
-
Lian, L. et al. Rapid volumetric bioprinting of decellularized extracellular matrix bioinks. Adv. Mater. 36, e2304846. https://doi.org/10.1002/adma.202304846 (2024).
-
Xie, M. et al. Volumetric additive manufacturing of pristine silk-based (bio)inks. Nat. Commun. 14, 210. https://doi.org/10.1038/s41467-023-35807-7 (2023).
-
Regehly, M. et al. Xolography for linear volumetric 3D printing. Nature 588, 620–624, https://doi.org/10.1038/s41586-020-3029-7 (2020).
-
Wolfel, A. et al. Bioxolography using diphenyliodonium chloride and N-vinylpyrrolidone enables rapid high-resolution volumetric 3D printing of spatially encoded living matter. Adv. Mater. e2501052, https://doi.org/10.1002/adma.202501052 (2025).
-
Stoecker, L. et al. Xolography for biomedical applications: dual-color light-sheet printing of hydrogels with local control over shape and stiffness. Adv. Mater. 37, e2410292. https://doi.org/10.1002/adma.202410292 (2025).
-
Habibi, M., Foroughi, S., Karamzadeh, V. & Packirisamy, M. Direct sound printing. Nat. Commun. 13, 1800. https://doi.org/10.1038/s41467-022-29395-1 (2022).
-
Derayatifar, M., Habibi, M., Bhat, R. & Packirisamy, M. Holographic direct sound printing. Nat. Commun. 15, 6691. https://doi.org/10.1038/s41467-024-50923-8 (2024).
-
Kuang, X. et al. Self-enhancing sono-inks enable deep-penetration acoustic volumetric printing. Science 382, 1148–1155, https://doi.org/10.1126/science.adi1563 (2023).
-
Wu, M. et al. Sound innovations for biofabrication and tissue engineering. Microsyst. Nanoeng. 10, 170, https://doi.org/10.1038/s41378-024-00759-5 (2024).
-
Vidler, C. et al. Dynamic interface printing. Nature 634, 1096–1102, https://doi.org/10.1038/s41586-024-08077-6 (2024).
-
Forsythe, S. D. et al. Environmental toxin screening using human-derived 3D bioengineered liver and cardiac organoids. Front. Public Health 6, 103, https://doi.org/10.3389/fpubh.2018.00103 (2018).
-
Wang, Z. et al. 3D bioprinting in cardiac tissue engineering. Theranostics 11, 7948–7969, https://doi.org/10.7150/thno.61621 (2021).
-
Motlagh, D., Yang, J., Lui, K. Y., Webb, A. R. & Ameer, G. A. Hemocompatibility evaluation of poly(glycerol-sebacate) in vitro for vascular tissue engineering. Biomaterials 27, 4315–4324, https://doi.org/10.1016/j.biomaterials.2006.04.010 (2006).
-
Fleischer, S., Tavakol, D. N. & Vunjak-Novakovic, G. From arteries to capillaries: approaches to engineering human vasculature. Adv. Funct. Mater. 30, 1910811. https://doi.org/10.1002/adfm.201910811 (2020).
-
Sexton, Z. A. et al. Rapid model-guided design of organ-scale synthetic vasculature for biomanufacturing. Science 388, 1198–1204, https://doi.org/10.1126/science.adj6152 (2025).
-
Oh, D., Kim, D. & Nam, S. Y. Predicting printability in suspended bioprinting using a rheology-informed hierarchical machine learning approach. Bioprinting 50, 00427, https://doi.org/10.1016/j.bprint.2025.00427 (2025).
-
Liao, H. et al. Data-driven de novo design of super-adhesive hydrogels. Nature 644, 89–95, https://doi.org/10.1038/s41586-025-09269-4 (2025).
-
Chen, S. L. et al. Machine learning-generated compression modulus database for 3D printing of gelatin methacryloyl. Int. J. Bioprinting 10, 560–573, https://doi.org/10.36922/ijb.3814 (2024).
-
Wu, Y., Ding, X., Wang, Y. & Ouyang, D. Harnessing the power of machine learning into tissue engineering: current progress and future prospects. Burns Trauma 12, tkae053. https://doi.org/10.1093/burnst/tkae053 (2024).
-
Kolesky, D. B., Homan, K. A., Skylar-Scott, M. A. & Lewis, J. A. Three-dimensional bioprinting of thick vascularized tissues. Proc. Natl. Acad. Sci. USA 113, 3179–3184, https://doi.org/10.1073/pnas.1521342113 (2016).
-
Hua, W. et al. High-speed embedded ink writing of anatomic-size organ constructs. Adv. Sci. 12, e2405980. https://doi.org/10.1002/advs.202405980 (2025).
-
Zhang, C. et al. Multiscale embedded printing of engineered human tissue and organ equivalents. Proc. Natl. Acad. Sci. USA 121, e2313464121. https://doi.org/10.1073/pnas.2313464121 (2024).
-
Zhao, J. Z., Hussain, M., Wang, M. N., Li, Z. Y. & He, N. Y. Embedded 3D printing of multi-internal surfaces of hydrogels. Addit. Manuf. 32, 101097. https://doi.org/10.1016/j.addma.2020.101097 (2020).
-
Ribezzi, D. et al. Shaping synthetic multicellular and complex multimaterial tissues via embedded extrusion-volumetric printing of microgels. Adv. Mater. 35, e2301673. https://doi.org/10.1002/adma.202301673 (2023).
-
Mancera Azamar, K. M. et al. Innovative engineering approaches to model host-microbiome interactions in vitro. Adv. Drug Deliv. Rev. 226, 115677. https://doi.org/10.1016/j.addr.2025.115677 (2025).
-
Kim, R. & Allbritton, N. L. A microphysiological system with an anaerobic air-liquid interface and functional mucus layer for coculture of intestinal bacteria and primary human colonic epithelium. Adv. Mater. Interfaces 11, https://doi.org/10.1002/admi.202400093 (2024).
-
Chen, Y. et al. Bioengineered 3D tissue model of intestine epithelium with oxygen gradients to sustain human gut microbiome. Adv. Health. Mater. 11, e2200447. https://doi.org/10.1002/adhm.202200447 (2022).
-
Cheng, L. et al. A 3D bioprinted gut anaerobic model for studying bacteria-host interactions. Research 6, 0058. https://doi.org/10.34133/research.0058 (2023).
-
Jalili-Firoozinezhad, S. et al. A complex human gut microbiome cultured in an anaerobic intestine-on-a-chip. Nat. Biomed. Eng. 3, 520–531, https://doi.org/10.1038/s41551-019-0397-0 (2019).
-
Tachedjian, G., Aldunate, M., Bradshaw, C. S. & Cone, R. A. The role of lactic acid production by probiotic Lactobacillus species in vaginal health. Res. Microbiol. 168, 782–792, https://doi.org/10.1016/j.resmic.2017.04.001 (2017).
-
Zheng, D. W. et al. A microbial community cultured in gradient hydrogel for investigating gut microbiome-drug interaction and guiding therapeutic decisions. Adv. Mater. 35, e2300977. https://doi.org/10.1002/adma.202300977 (2023).
-
Ahmed, M. & Ffrench-Constant, C. Extracellular matrix regulation of stem cell behavior. Curr. Stem Cell Rep. 2, 197–206, https://doi.org/10.1007/s40778-016-0056-2 (2016).
-
Singh, B., Fleury, C., Jalalvand, F. & Riesbeck, K. Human pathogens utilize host extracellular matrix proteins laminin and collagen for adhesion and invasion of the host. FEMS Microbiol. Rev. 36, 1122–1180, https://doi.org/10.1111/j.1574-6976.2012.00340.x (2012).
-
Muniraj, G. et al. Microphysiological modeling of gingival tissues and host-material interactions using Gingiva-on-chip. Adv. Health. Mater. 12, e2301472. https://doi.org/10.1002/adhm.202301472 (2023).
-
Schwartzentruber, J. et al. Molecular and functional variation in iPSC-derived sensory neurons. Nat. Genet. 50, 54–61, https://doi.org/10.1038/s41588-017-0005-8 (2018).
-
Zhang, J. et al. An immune-competent human gut microphysiological system enables inflammation-modulation by Faecalibacterium prausnitzii. NPJ Biofilms Microbiomes 10, 31. https://doi.org/10.1038/s41522-024-00501-z (2024).
-
Katholnig, K., Linke, M., Pham, H., Hengstschlager, M. & Weichhart, T. Immune responses of macrophages and dendritic cells regulated by mTOR signalling. Biochem. Soc. Trans. 41, 927–933, https://doi.org/10.1042/BST20130032 (2013).
-
d’Hennezel, E., Abubucker, S., Murphy, L. O. & Cullen, T. W. Total lipopolysaccharide from the human gut microbiome silences toll-like receptor signaling. mSystems 2, https://doi.org/10.1128/mSystems.00046-17 (2017).
-
Kim, M.-H., Kim, D. & Sung, J. H. A Gut-Brain Axis-on-a-Chip for studying transport across epithelial and endothelial barriers. J. Ind. Eng. Chem. 101, 126–134, https://doi.org/10.1016/j.jiec.2021.06.021 (2021).
-
Sarah, R. et al. Data-driven optimization of bioink formulations for extrusion-based bioprinting: a predictive modeling approach. J. Manuf. Sci. Eng. 147, https://doi.org/10.1115/1.4069041 (2025).
-
Xu, Y., Sarah, R., Habib, A., Liu, Y. & Khoda, B. Constraint based Bayesian optimization of bioink precursor: a machine learning framework. Biofabrication 16, https://doi.org/10.1088/1758-5090/ad716e (2024).
-
Rasouli, R., Sweeney, C. & Frampton, J. P. Heterogeneous and composite bioinks for 3D-bioprinting of complex tissue. Biomed. Mater. Devices 3, 108–126, https://doi.org/10.1007/s44174-024-00171-7 (2025).
-
Lee, H. & Cho, D. W. One-step fabrication of an organ-on-a-chip with spatial heterogeneity using a 3D bioprinting technology. Lab Chip 16, 2618–2625, https://doi.org/10.1039/c6lc00450d (2016).
-
Florczak, S. et al. Adaptive and context-aware volumetric printing. Nature 645, 108–114, https://doi.org/10.1038/s41586-025-09436-7 (2025).
-
Maharjan, S. et al. 3D bioprinting of plant and animal cell-based hybrid food. Nat. Commun. 16, 6935. https://doi.org/10.1038/s41467-025-61996-4 (2025).
Acknowledgements
All figures were organized in Adobe Illustrator; icons are borrowed from BioRender.com. The authors used AI-based language editing tools solely to improve grammar, readability, and clarity. All scientific content and conclusions were developed, written, and verified by the authors.
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Hua, W., Gaharwar, A.K. 3D biofabricated in vitro models as new approach methodologies for animal alternatives. npj Biomed. Innov. 3, 20 (2026). https://doi.org/10.1038/s44385-026-00073-x
-
Received:
-
Accepted:
-
Published:
-
Version of record:
-
DOI: https://doi.org/10.1038/s44385-026-00073-x