Researchers have discovered how “pirate phages” hijack other viruses to break into bacteria, sharing new genetic material for dangerous traits.
Imperial scientists have uncovered how bacteriophages are able to hijack other viruses to break into bacterial cells and spread, through an act of microbial piracy which could potentially be harnessed for medicine.
The discovery, published in the journal Cell, reveals a major route by which bacteria are able to acquire new genetic material, including traits that can make them more virulent or more resistant to antibiotics. The researchers believe it could also open the door to new ways of tackling the global threat of antimicrobial resistance (AMR) and developing rapid diagnostic tools.
Phages (or bacteriophages) are viruses that infect and kill bacteria. They are among the most abundant organisms on Earth and are often highly specific, each tailored to attack just one bacterial species. Structurally, they resemble microscopic syringes: with a “head” section packed with DNA and a tail section tipped with spiky fibers that latch onto bacteria and inject their genetic payload.
But phages themselves are not safe from parasites. They can be targeted by small genetic elements known as phage satellites that hijack the phage’s own genetic machinery to propagate.
In the latest study, Imperial researchers focused on a powerful family of phage satellites called capsid-forming phage-inducible chromosomal islands (cf-PICIs). These genetic elements can spread genes for antibiotic resistance and virulence, and are found across more than 200 bacterial species. Exactly how they managed to move so efficiently, however, was unclear.
Microbial piracy
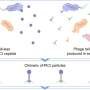
First discovered by the team in 2023, cf-PICIs can build their own capsids (the viral “heads”), but they lack tails, meaning on their own they produce non-infective particles—i.e., they are not able to infect phages. In this latest work, researchers at Imperial’s Center for Bacterial Resistance Biology discovered the missing piece of the puzzle: cf-PICIs hijack tails from unrelated phages, creating hybrid “chimeric” viruses. The result is a chimeric phage carrying cf-PICI DNA inside their own capsids but a phage-derived tail attached.
Crucially, some cf-PICIs can hijack tails from entirely different phage species, effectively broadening their host range. Because the tail decides which bacteria are targeted, this piracy gives cf-PICIs the ability to infiltrate new bacterial species, explaining their great abundance in nature.
According to the researchers, the implications could be important for science. By understanding and harnessing this molecular piracy, researchers believe they could re-engineer satellites to target antibiotic-resistant bacteria, overcome stubborn bacterial defenses such as biofilms, and even develop powerful new diagnostic tools.
“These pirate satellites don’t just teach us how bacteria share dangerous traits,” explains Dr. Tiago Dias da Costa, from Imperial’s Department of Life Sciences. “They could inspire next-generation therapies and tests to outmaneuver some of the most difficult infections we face.”
The Imperial team has successfully filed patents to further develop the work and hopes to begin testing the translational applications of the technology.
Professor Jose Penades, from Imperial’s Department of Infectious Disease, said, “Our early work first identified these odd genetic elements, where we found they are effectively a parasite of a parasite. We now know these mobile genetic elements form capsids which can swap ‘tails’ taken from other phages to get their own DNA into a host cell. It’s an ingenious quirk of evolutionary biology, but it also teaches us more about how genes for antibiotic resistance can be spread through a process called transduction.”
Dr. Dias da Costa, added, “This experimental work sheds more light on a crucial method of gene transfer in bacteria. If we can harness and engineer cf-PICIs it could provide us with a valuable new tool in the fight against antimicrobial resistance.”
AI co-scientist tool
In a linked project, coordinated through the Fleming Initiative—a partnership between Imperial College London and Imperial College Healthcare NHS Trust—researchers used their experimental work to validate a groundbreaking AI platform developed by Google.
Dubbed the “co-scientist,” the platform is designed to help scientists develop smarter experiments and accelerate discovery. This research is also published in Cell.
To test the platform, the Imperial team posed the same basic scientific questions that had driven their own work: How do cf-PICIs spread across so many bacterial species?
Armed with this starting point, and drawing on web searches, research papers, and databases, the AI independently generated hypotheses that mirrored the team’s own experimentally proven ideas—effectively pointing to the same experiments that had taken years of work to establish, but doing so in a matter of days.
The researchers say this shows the extraordinary potential of AI systems to “super-charge science,” not by replacing human insight, but by accelerating it. They are now working with Google to further develop the platform and explore how it could transform the pace of biomedical research.
More information: Lingchen He et al, Chimeric infective particles expand species boundaries in phage-inducible chromosomal island mobilization, Cell (2025). DOI: 10.1016/j.cell.2025.08.019
José R. Penadés et al, AI mirrors experimental science to uncover a mechanism of gene transfer crucial to bacterial evolution, Cell (2025). DOI: 10.1016/j.cell.2025.08.018
Journal information: Cell
Citation: ‘Microbial piracy’ uncovers new way to fight drug-resistant infections (2025, September 9) retrieved 9 September 2025 from https://phys.org/news/2025-09-microbial-piracy-uncovers-drug-resistant.html
This document is subject to copyright. Apart from any fair dealing for the purpose of private study or research, no part may be reproduced without the written permission. The content is provided for information purposes only.