Abstract
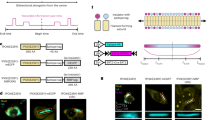
Cells constantly change their molecular state in response to internal and external cues1. Mapping cellular activity in tissues with spatiotemporal precision is essential for understanding organ physiology, pathology, and regenerative processes. Current cell-sensing modalities primarily rely on either endpoint analysis that takes static snapshots, or real-time sensing that monitors a small subset of cells3,4. Here, we introduce Granularly Expanding Memory for Intracellular Narrative Integration (GEMINI), an in cellulo recording platform that leverages a computationally designed protein assembly as an intracellular memory device to record the history of individual cells. GEMINI grows predictably within live cells, capturing cellular events as tree-ring-like fluorescent patterns for imaging-based retrospective readout. Absolute chronological information of activity histories is attainable with hour-level accuracy. GEMINI effectively maps differential NFκB-mediated transcriptional changes, resolving fast dynamics of 15 minutes and providing quantifiable signal amplitudes. In a xenograft model, GEMINI records inflammation-induced signaling dynamics across tissue, revealing spatial heterogeneity linked to vascular density. When expressed in the mouse brain, GEMINI minimally impacts neuronal functions and can resolve both transcriptional changes and activity patterns of neurons. Together, GEMINI provides a robust and generalizable means for spatiotemporal mapping of cell dynamics underlying physiological and pathological processes in both culture and intact tissues.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Rent or buy this article
Prices vary by article type
from$1.95
to$39.95
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
Supplementary information
Supplementary Information
This file contains additional notes about GEMINI growth modelling, Supplementary Methods, Supplementary Figs. 1–15, Supplementary Tables 1–2, and Supplementary References.
Reporting Summary
Supplementary Data
Source data for Figs. 4 and 5, Extended Data Figs. 7-10, and Supplementary Figs. 14 and 15.
Peer Review File
Supplementary Video 1
GEMINI growth in HEK293T cells. Time-lapse video showing GEMINI growth in the HEK293T GEMINI clonal cell line.
Supplementary Video 2
Tracking the growth of single GEMINI particles. Time-lapse videos with single-particle tracking (left) and the resolved growth profiles (right) of GEMINI particles.
Rights and permissions
About this article
Cite this article
Yan, Y., Lu, J., Li, Z. et al. Genetically encoded assembly recorder temporally resolves cellular history. Nature (2026). https://doi.org/10.1038/s41586-026-10323-y
-
Received:
-
Accepted:
-
Published:
-
DOI: https://doi.org/10.1038/s41586-026-10323-y