Main
The modern chemical industry relies on the consumption of finite fossil resources at a pace and scale that is inherently unsustainable. These processes are energy intensive and generate consumer products that are ultimately disposed by landfill or incineration, resulting in the irretrievable loss of this valuable carbon as environmental pollution or CO2 in the atmosphere. By contrast, nature has evolved elegant and efficient mechanisms for carbon resource utilization, by-product recycling and sustainable chemical synthesis. As these processes are genetically encoded, they offer a blueprint for modern engineering biology to remediate and upcycle carbon embedded in industrial and post-consumer waste. The resulting engineered bioprocesses can reintegrate this carbon into the circular chemical economy, while simultaneously reducing pollution, greenhouse gas emissions and the underlying drivers of global climate change.
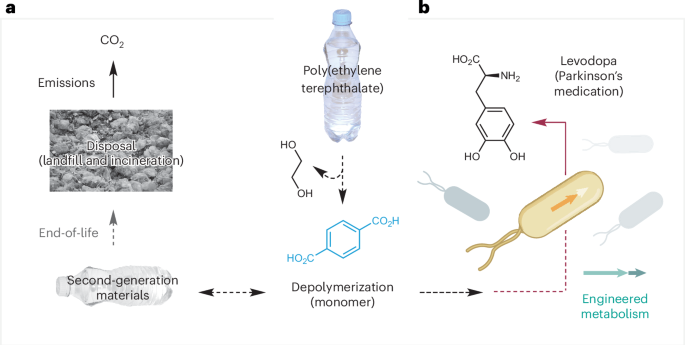
Research in this area has focussed on the bioavailable polymers cellulose, chitin and lignin as substrates, but more recently has explored the use of plastic waste as a microbial feedstock. Enzymatic depolymerization of lignocellulose and chitin has yielded carbohydrate monomers for bacterial growth and upcycling pathways to biofuels1, polyhydroxyalkanoates2, terpenes3 and amino acids4, whereas aromatic monomers isolated from lignin have been used as cellular substrates for adipic acid5 and coniferyl alcohol6 synthesis. The bio-upcycling of plastic waste has been enabled by the discovery of enzymes capable of depolymerizing poly(ethylene terephthalate) (PET)7, catalysed by the discovery of Ideonella sakaiensis, a bacterium capable of depolymerizing and assimilating PET8. This breakthrough has since driven the engineering of IsPETase variants with enhanced catalytic performance for applications in industrial biotechnology (Fig. 1a). These efforts have yielded PET hydrolases with enhanced activity and thermal stability9,10, as well as metabolic pathways for upcycling PET-derived monomers into value-added products such as vanillin11, adipic acid12, paracetamol13 and other platform chemicals14,15. In addition, there exists a growing list of chemoenzymatic approaches to upcycle other plastic wastes such as poly(hydroxybutyrate) into acetone16, polyethylene into functionalized carboxylic acids17 and mixed plastic wastes into ß-ketoadipate and poyhydroxyalkanoates18. Chemical catalysts have also been interfaced with plastic upcycling pathways to create new biocompatible chemistry approaches and green synthetic chemistry methods19 for the mild conversion of a range of plastic wastes into industrially applicable second-generation chemical products. Of note, ref. 20 recently reported the use of chemical catalysis to convert post-consumer polystyrene to benzoic acid (Co(NO3)2 + Mn(NO3)2, N-hydroxyphthalimide, O2 (4 bar), acetic acid, 150 °C followed by microbial conversion to the complex small molecules ergothioneine, mutilin and pleuromutilin in engineered Aspergillus nidulans and as a sole carbon source to generate the atoxigenic biocontrol agent Aspergillus flavus Af36. Similarly, another study18 coupled the chemocatalysed autooxidation of mixed domestic plastic wastes to oxygenated carboxylic acid intermediates with biotransformation by engineered strains of Pseudomonas putida KT2440 to create a metabolic funnel to valuable synthetic building blocks. Inspired by recent advances and the emerging potential of engineering biology to upcycle plastic waste into high-value molecules, we report the microbial conversion of industrial PET waste and a post-consumer PET plastic bottle into levodopa (l-DOPA), a frontline treatment for the symptoms of Parkinson’s disease (Fig. 1b). l-DOPA is currently produced at a global scale of ~250 t yr−1 (ref. 21) with demand projected to rise as a result of increased disease prevalence22,23. Beyond its clinical importance, l-DOPA serves as a biosynthetic precursor to melanin and a variety of complex plant-derived natural products24,25. Despite the existence of several biotechnological routes to l-DOPA—including fermentative pathways from l-tyrosine, chitin or D-glucose and expression in transgenic tomatoes—commercial production remains reliant on fossil fuel-derived chemical or chemoenzymatic synthesis26,27,28,29,30. Each alternative faces notable challenges, such as poor carbon economy, oxidative degradation, inefficient feedstock utilization or regulatory hurdles. This work therefore offers a sustainable alternative to existing chemical and biological methods for producing l-DOPA from virgin petrochemicals and highlights the first application of engineering biology to valorize plastic waste into a therapeutic for neurological disease.
Approaches to the recycling, upcycling and environmental disposal of PET plastic waste, including the proposed bio-upcycling of PET waste to the Parkinson’s medication l-DOPA in engineered bacteria. a, Current: closed-loop recycling. b, This work: microbial upcycling. Credit: photographs in a, Rawpixel (https://www.rawpixel.com); bacterial icon in b, Bioicons (https://bioicons.com).
Results
De novo pathway design and optimization
l-DOPA synthesis from PET monomer terephthalic acid (TPA) was envisioned via a four-step biosynthetic pathway encoded by seven genes in the laboratory bacterium Escherichia coli BL21(DE3). The pathway proceeds via conversion of TPA to protocatechuate (PCA) catalysed by a terephthalate 1,2-dioxygenase complex (TphA2 and TphA3), and cognate reductase (TphA1; together referred as the TPADO complex) and dihydroxy-3,5-cyclohexadiene-1,4-dicarboxylic acid dehydrogenase (known as DCDDH or TphB) from Comamonas sp. Decarboxylation of PCA to catechol by AroY and prFMN cofactor regeneration enzyme KpdB from Klebsiella pneumoniae is then followed by a final C–C bond formation via electrophilic aromatic substitution between catechol and pyruvate in the presence of ammonia by the pyridoxal 5′-phosphate (PLP)-dependent tyrosine-phenol lyase (TPL) from Fusobacterium nucleatum to form l-DOPA (Fig. 2a). The tpado and dcddh genes were assembled on to a pGro7 derived backbone to generate pPCA1 (module 1), aroY and kpdB genes were assembled on to a pQLinkN vector to generate pCAT1 (module 2) and tpl was assembled in a joint universal modular plasmids (JUMP) assembly level 1 vector to generate pFnTPL (module 3). The activity of each module was examined by transforming E. coli BL21(DE3) cells with pPCA1, pCAT1, pPCA1 + pCAT1 or pFnTPL, confirming soluble protein expression after growth in LB medium by SDS–PAGE (Supplementary Figs. 1 and 2) and then incubating with the requisite pathway substrate or intermediate in a whole-cell biotransformation (optical density OD600 = 30).
a, De novo biosynthetic pathway to l-DOPA from PET monomer TPA. b, Whole-cell activity when pPCA1, pCAT1 and pFnTPL are singly and multiply expressed in E. coli BL21(DE3) as well as whole pathway with pPCA1_pCAT-FnTPL. c, Enhanced TPA conversion to PCA in strains E. coli_pPCA1 and E. coli_pPCA3 expressing tpaK from R. jostii. d, PCA and catechol conversions upon increasing terephthalate concentration in strains E. coli_pPCA3 and E. coli_pPCA3_pCAT1, respectively. Metabolite concentrations were determined by reverse-phase HPLC relative to an internal standard of caffeine (0.01 g l−1). Data are presented as an average of three replicate experiments to one standard deviation. All data shown are from 3 ml reactions performed in 15 ml Falcon tubes. In b, the supercripts are as follows: [a]TPA substrate; [b]PCA substrate; and [c]catechol substrate. Credit: photograph in a, Rawpixel (https://www.rawpixel.com); bacterial icons in a, Bioicons (https://bioicons.com).
Cells containing TPADO-dcddh (module 1), AroY/KpdB (module 2) or TPL (module 3) enzymes were highly active in isolation and observed to convert >90% substrate to product in 24 h (Fig. 2b). We sought to improve diffusion of terephthalate across the negatively charged bacterial cell membrane. This is known to be most efficient at pH 5 (ref. 11) (pKa1 = 3.5; pKa2 = 4.3) and therefore to mitigate this at pH 7 we assembled tpaK from Rhodococcus jostii downstream of the tpado and dcddh genes via homologous end assembly to generate the modified plasmid pPCA3 (Supplementary Fig. 3). TpaK is part of the major facilitator superfamily class of membrane transporters and has been shown to import aromatic acids including TPA when heterologously expressed in P. putida31,32. Pleasingly, TPA conversion to PCA at pH 7 displayed accelerated product formation (Fig. 2c). To our knowledge this constitutes the first use of a TPA transporter for biocatalysis in E. coli.
Co-transformation with pCAT1 generated the modified strain E. coli_pPCA3_pCAT1 for conversion of TPA to catechol. Despite high-level production of PCA by E. coli_pPCA3, conversion to catechol was efficient at low concentration (1–10 mM; >85%) but decreased to <40% with increasing TPA loading (Fig. 2d). Intriguingly, the addition of 30 mM catechol also resulted in a dramatic reduction in conversion of TPA to PCA in this strain, indicating that catechol inhibits TPADO activity at this concentration (Supplementary Fig. 5). Although the precise mechanism of inhibition is unclear, preliminary molecular docking simulations indicated favourable binding interactions between catechol and TPADO in and around the active site and ortho-quinone formation under the oxygenic conditions required for TPA conversion to PCA could covalently inhibit TPADO or compete for Fe binding within the active site. Additionally, catechol is known to inactivate proteins through cross-linking and has been shown to inhibit reductases33,34. Regardless, it was hypothesized that catechol mediated TPADO inhibition would be limited through conversion to l-DOPA within the full biosynthetic pathway.
Pathway inhibition and in silico modelling
To circumvent the accumulation of catechol in whole-cell reactions containing TPADO, E. coli_pPCA1 was transformed with pCAT-FnTPL encoding the highly active TPL from F. nucleatum, generating the fully engineered strain E. coli_pPCA1_pCAT-FnTPL for TPA to l-DOPA biosynthesis. However, incubation of E. coli_pPCA1_pCAT-FnTPL in the presence of 5 mM TPA produced 0.99 mM catechol (20%) and no detectable l-DOPA (Fig. 2b). We hypothesized that TPL was being inhibited by upstream pathway intermediates during l-DOPA biosynthesis. Indeed, conversion of catechol to l-DOPA in E. coli_pFnTPL reduced from 80% to 0% when catechol was incubated in the presence of >1 mM PCA (Fig. 3d and Supplementary Fig. 6). Similar results were observed in vitro using His-tag purified FnTPL, with no l-DOPA produced at >2 mM PCA (Supplementary Fig. 6). Overall, this provided evidence to suggest the PCA-dependent inhibition of TPL during l-DOPA biosynthesis. To support this hypothesis, molecular docking simulations of catechol and PCA within the active site of FnTPL were performed using GNINA (Fig. 3). Key hydrogen bonding and electrostatic interactions were identified between PCA and conserved active site residues T127, K260, R384 and Y74. Free energy calculations showed that both PCA and catechol bind with identical and thermodynamically favourable energies (−5.99 kcal mol−1), suggesting that competitive binding probably occurs within the active site pocket. This aligns with recent reports of TPL inhibition by gallic acid and 3,5-dihydroxybenzoic acid in bacteria associated with mouse gut microbiota35. Together, these findings provide strong evidence of PCA-dependent TPL inhibition during l-DOPA biosynthesis and highlight the need to circumvent this bottleneck to enable efficient l-DOPA production from TPA in vivo.
Predicted binding modes of PCA or catechol with FnTPL–dimer–PLP complex. a, Model of the functional dimer of FnTPL, with covalently linked PLP and docked catechol in monomer A (light grey) shown as stick representation in the active site. b, Predicted binding of catechol in the active site mediated primarily by phenylalanine and arginine residues. The catalytic Tyr-74 contributed by monomer B is highlighted in dark grey. c, Docked PCA bound to FnTPL with interactions mediated by arginine residues augmented by Met-382, Thr-127 and Phe-120. Both substrates are predicted to have a favourable binding affinity of −5.99 kcal mol−1, with Arg-220, Arg-384, Arg-407 and Thr-52 contributing to the binding of both substrates. Molecular docking simulations were performed using the molecular docking package GNINA. d, Assaying pathway module inhibition by pathway intermediates using E. coli_pFnTPL cells incubated in the presence of catechol and varying concentrations of PCA. Data are presented as an average of three replicate experiments to one standard deviation.
To this end, the three pathway modules were separated into two strains (E. coli_pPCA3_pCAT1 and E. coli_pFnTPL) to enable catechol accumulation before conversion to l-DOPA. Reaction optimizations to maximize the performance of module 3 within E. coli_pFnTPL identified pH, time and pyruvate concentration as critical to high l-DOPA conversion from exogenous catechol in this strain (Fig. 4b). As reported in the literature36, FnTPL was most efficient at pH 8 (Fig. 4b). Interestingly, l-DOPA concentrations also decreased by 40% from 4.2 mM to 2.5 mM over 17 h under these optimized reaction conditions. Although l-DOPA is known to polymerize via dopaquinone to poly(dopaquinone) under aerobic conditions37, the addition of antioxidants did not reverse the observed product loss over time. Liquid chromatography–mass spectrometry (LC-MS) analysis of reaction extracts instead confirmed the degradation of l-DOPA via a non-enzymatic Pictet–Spengler reaction with pyruvate to form a heterocyclic adduct (Supplementary Fig. 7)27. Together, the optimum reaction conditions for l-DOPA synthesis from 5 mM catechol by E. coli_pFnTPL at OD600 = 30 were concluded to be pH 8.0 in the presence of 60 mM pyruvate for 3 h at 21 °C (Fig. 4b).
a, A one-pot two-strain approach to decouple PCA mediated inhibition of TPL for l-DOPA biosynthesis from TPA and PET. b, Whole-cell reaction optimization for conversion of exogenous catechol to l-DOPA by E. coli_pFnTPL. c, Time course of two strain system for conversion of TPA to l-DOPA, with 24 h of reaction from TPA to CAT by strain E. coli_pPCA3_pCAT1 before addition of strain E. coli_pFnTPL for 24 h. A scaling factor of 0.83 was applied to account for dilution effects from the addition of two strains. d, Capture of released CO2 from TPA whole-cell reactions in E. coli by C. reinhardtii CC1690. Peak areas of residual CO2 for C. reinhardtii grown on TAP media (control) compared with incubation with or without headspace gas mixture from E. coli (HSGM). e, Preparative microbial biosynthesis of l-DOPA salt from industrial HSF PET waste collected from API Foilmakers. Data are presented as an average of three replicate experiments to one standard deviation, except e which was performed as a scaled-up single replicate. Catechol was generated by strain E. coli_pPCA3_pCAT1. ND, not detected. Credit: photographs in a, Rawpixel (https://www.rawpixel.com); Chlamydomonas icon in d, DBCLS/TogoTV under a Creative Commons license CC BY 4.0.
Two-strain bioconversion
Having optimized conditions for the conversion of TPA to catechol by E. coli_pPCA3_pCAT1 and catechol to l-DOPA by E. coli_pFnTPL, we moved on to combine these strains for the one-pot biotransformation of TPA to l-DOPA. As PCA-dependent inhibition of TPL had been observed at concentrations >250 µM, a reaction involving the sequential addition of E. coli_pFnTPL cells after incubation of E. coli_pPCA3_pCAT1 with TPA was envisaged. Additionally, analysis of E. coli_pPCA3_pCAT1 (OD600 = 30) cells incubated with 5 mM TPA at 21 °C (220 rpm) for 24 h showed >90% catechol was present in the cell supernatant, indicating this would be suitable for uptake by a second microorganism (Supplementary Fig. 8). To this end, TPA was incubated with E. coli_pPCA3_pCAT1 (OD600 = 30) at 21 °C (220 rpm) for 24 h, before the addition of E. coli_pFnTPL (OD600 = 30) and further incubation for 3 h. Pleasingly, this resulted in the production of l-DOPA from TPA in 0.68 g l−1 and 69% overall conversion as the major product by high-performance liquid chromatography (HPLC) (Fig. 4c and Supplementary Fig. 9).
CO2 capture using microalgae
As a proof-of-concept and preliminary assessment of whether CO2 released during the enzymatic decarboxylation of TPA to catechol could be offset, we tested whether the microalga Chlamydomonas reinhardtii CC1690 could recapture this CO2 via photosynthesis. Engineered E. coli cultures supplied with 5 mM TPA-generated elevated CO2 levels (Supplementary Fig. 10) and headspace gas from these cultures was transferred to actively growing C. reinhardtii. Within 12 h, CO2 levels fell below the detection limit only in algal cultures and these cultures showed increased optical density and chlorophyll content relative to controls, confirming both CO2 fixation and stimulated algal growth (Fig. 4d and Supplementary Figs. 11 and 12). While these preliminary findings suggest that TPA-derived CO2 can be assimilated into algal biomass, further development, quantitative analysis and system-level validation will be required to establish the extent to which coupling biosynthesis with microalgal CO2 capture can contribute to overall process carbon neutrality.
Industrial waste valorization and product isolation
Following on from the one-pot bioconversion of PET monomer TPA to l-DOPA, we next moved on to generate l-DOPA from industrial and post-consumer PET waste. In addition to the use of a post-consumer PET bottle and packaging, we examined the upcycling of industrial PET waste. Hot stamping foils (HSF) are a prolific source of plastic waste worldwide generated by the chemical industry from the depositing of ultrathin lacquer and adhesive labels. This industry is rapidly growing (US$2.9 billion market in 2022, 5.6% CAGR through 2032) and is estimated to generate 40,000 t of PET waste per annum globally (https://www.maximizemarketresearch.com/market-report/global-hot-stamping-foils-market/25706). We depolymerized a PET bottle (from discarded waste at the University of Edinburgh, UK) and HSF samples (from API Foilmakers) under alkaline conditions, generating TPA-containing product streams that were quantified by nuclear magnetic resonance spectroscopy (51% and 83% purity for PET bottle and HSF waste, respectively). Crude TPA-rich samples were then added to optimized one-pot reactions using strains E. coli_pPCA3_pCAT1 (pH 7, 24 h, 21 °C, 220 rpm) and E. coli_pFnTPL (pH 8, 3 h, 21 °C, 220 rpm). Pleasingly, l-DOPA was generated from all reactions at 2.0 mM (49%) and 2.3 mM (55%) conversion from bottle and stamping foil PET waste, respectively (Supplementary Fig. 13). Furthermore, enzymatic depolymerization of PET packaging film by LCCICCM generated 4.64 mM l-DOPA from the released TPA (Supplementary Fig. 14). Reduced product conversions were attributed to the presence of lower grade PET from post-consumer waste due to residual plasticizers. The reaction was then scaled-up to 0.5 l and yielded 0.9 g l−1 of l-DOPA in one-pot process directly from chemically depolymerized PET waste (Supplementary Fig. 13).
We next sought to isolate l-DOPA from biotransformation reactions using preparative reverse-phase HPLC. For efficient chromatographic separation, the reactions were miniaturized to generate low-volume product streams with high l-DOPA concentrations. This was achieved using a redesigned plasmid, pPCA4, which encodes tpado, dcddh and tpaK genes on a pBR322/rop origin backbone, compared with the p15a origin in pPCA3. This modification led to markedly improved catechol production at 30 mM TPA loading, increasing from 12.6 mM with pPCA3 to 27.6 mM with pPCA4 (Supplementary Fig. 15). In a two-step reaction involving E. coli_pPCA4_pCAT1 and E. coli_pFnTPL, we achieved 25.3 mM l-DOPA (5.0 g l−1, 84% conversion) from 0.26 g of stamping foil-derived TPA following sequential incubations of 24 h (strain 1) and 3 h (strain 1 + 2) at 21 °C (Fig. 4e). l-DOPA was successfully isolated as a solid TFA salt via preparative reverse-phase HPLC, equivalent to several clinical doses typically prescribed for early-onset Parkinson’s disease.
Discussion
We acknowledge that global plastic waste generation (~100 million t annually) far exceeds pharmaceutical production volumes. This pathway is therefore not proposed as a standalone solution, but rather as one component of a broader bio-upcycling portfolio. This study specifically focuses on stamping foil waste—a specialized and currently underaddressed industrial PET stream—as a viable feedstock for sustainable chemical manufacturing.
Further optimization towards industrial implementation will require pathway intensification to enable l-DOPA recovery via direct precipitation from fermentation broth, given its aqueous solubility (<5 g l−1 at 20 °C)26. Additional analysis will also be necessary to confirm the absence of plasticizers and other contaminants originating from stamping foil waste in the final product stream. In parallel, genomic integration of the de novo pathway will be essential to eliminate reliance on antibiotic selection at scale. This study serves as a proof-of-concept demonstration of feasibility rather than a fully optimized production process.
To enhance process sustainability, glucose recovered from surplus bread waste was shown to support biotransformation with no loss in efficiency (Supplementary Fig. 16). In addition, photosynthetic CO2 capture by C. reinhardtii presents a promising route towards carbon-neutral operation, although further development is needed to fully realize this potential (Fig. 4d). Future work will focus on life-cycle and technoeconomic assessments during bioreactor-scale intensification to rigorously quantify the environmental and economic benefits of the process, as well as extending the pathway towards the biosynthesis of more complex, medicinally relevant alkaloids from plastic waste using newly engineered microbial systems.
The conversion of waste carbon into high-value industrial chemicals remains a key objective for advancing the circular chemical and bioeconomy. This study explores the microbial transformation of TPA sourced from industrial and post-consumer PET waste into l-DOPA, a frontline medication for Parkinson’s disease. A de novo biosynthetic pathway was constructed in E. coli, addressing design challenges related to monomer import and feedback inhibition trough heterologous transporter expression and distribution across two cooperative strains. The resulting bioprocess operates under mild, aqueous conditions (pH 7–8), achieving high titres (5.0 g l−1) and enabling the isolation of preparative quantities of l-DOPA from both industrial PET waste and a single post-consumer plastic bottle. Notably, the aromatic structure of the PET monomer is preserved throughout biosynthesis and embedded in the final pharmaceutical product, eliminating the need for virgin fossil carbon. Overall, this work establishes engineering biology as a route to recover and repurpose carbon from plastic waste otherwise destined for landfill, incineration or environmental pollution into high-value therapeutics for neurodegenerative disease.
Methods
Detailed methods and protocols for all experiments reported in this study are provided in Supplementary Information.
Strains and plasmids
All cloning procedures were performed in E. coli DH5α. Genes encoding tphA1A2A3, dcddh, aroY and kpdB were synthesized in our previous reports11,12. The tpaK and tpl genes were codon-optimized for E. coli BL21(DE3) and synthesized using GeneArtTM (Thermo Scientific) and His-tagged tpl (hisfntpl) used the tpl sequence with additional was synthesized by Integrated DNA Technologies (Supplementary Table 3). The pET21a plasmid containing lcc was acquired as a gift from the laboratory of C.U.
All other recombinant plasmids were constructed by homology-mediated DNA assembly or by modular cloning. Homology-mediated DNA assembly was performed using the NEBuilder HiFi DNA Assembly Cloning and modular cloning was conducted by following the JUMP protocol38. Plasmids used in this study are listed in Supplementary Table 4 and were confirmed by colony PCR and Sanger sequencing (Azenta).
Oligonucleotide primers for generating plasmid pPCA3 from pPCA1 and pTpaK by homology-mediated DNA assembly were synthesized by Integrated DNA Technologies, these are pTpaK-F (tcgctgctttgagacgtactagtagcggccagctttaatacgactcactataggggaattgtgagcgga) and pTpaK-R (cataatacctagggctgagctagccgtaaaagcttatcgatgataagctgtcaaacatgagaattacaac).
Protein expression using E. coli BL21(DE3)
A single colony of E. coli BL21(DE3) cells transformed with appropriate expression plasmids was used to inoculate a 10 ml LB starter culture in a 50 ml Falcon tube containing the appropriate antibiotics and incubated overnight. For protein expression, the starter culture was back diluted to OD600 = 0.1 in 200 ml of LB and incubated in a 500 ml Erlenmeyer flask until the OD600 reached 0.6–0.8. At this point, isopropyl β-d-1-thiogalactopyranoside was added (final concentration of 0.5 mM) and cultured were incubated at 21 °C and 220 rpm for all proteins except FnTPL that was incubated at 30 °C and 220 rpm for 24 h before harvesting cells by centrifugation at 3,000g, 4 °C for whole-cell biotransformations or purification.
Whole-cell biotransformations
Harvested post-expression E. coli BL21(DE3) were washed with PBS, resuspended at OD600 = 30 in M9 medium and 3 ml of the resulting cell suspension was used for each biotransformation. For TPA to PCA and PCA to CAT, disodium terephthalate or PCA (final concentration of 5 mM) and glucose (final concentration of 5% w/v, unless stated otherwise) were added and incubated in a 15 ml Falcon tube at 21 °C and 220 rpm for 24 h. For CAT to l-DOPA, CAT (final concentration of 5 mM), sodium pyruvate (60 mM), ammonium chloride (250 mM), potassium chloride (1 mM), disodium EDTA (3 mM), sodium sulfite (7 mM) and pyridoxal phosphate (1 mM) were added and incubated in a 15 ml Falcon tube at 21 °C and 220 rpm for 3 h. The supernatant was analysed by HPLC (Supplementary Table 1).
Sequestration of TPA-derived CO2 by C. reinhardtii
E. coli_pPCA4_pCAT1 cells were resuspended in M9-glucose (2%) at OD600 = 30 and 2.85 ml was aliquoted into headspace vials and made up to 3 ml with either 100 mM disodium terephthalate stock (5 mM final concentration) or sterile double-distilled H2O. Vials were immediately sealed with a crimp lid and incubated (21 °C, 220 rpm, 24 h). Pregrown C. reinhardtii CC1690 culture in tris-acetate-phospate (TAP) medium was aliquoted into sterile transparent glass headspace vials at a volume of 5 ml and immediately sealed with a crimp lid. To enable the transfer of evolved CO2 from E. coli cultures to C. reinhardtii cultures or TAP control vials, E. coli samples were heated to 70 °C in a water bath (10 min) to increase gas abundance in the headspace. Evolved gas (15 ml) was then drawn into a rubber-stoppered syringe through a 0.22 μm PTFE filter using a 27-gauge needle then sealed using a Luer Lock (Agilent). The needle was then sterilized by heating under a flame then evolved gas was injected into experimental samples. Pressure was applied to the septum of the crimp lid as needle was removed to promote resealing then samples were incubated under continuous illumination as previously described for desired timeframe. Gases within E. coli, TAP and C. reinhardtii samples were identified at desired timepoints by headspace gas chromatography as detailed in Supplementary Information and catechol production was analysed by HPLC.
Preparative biotransformation of PET waste TPA to l-DOPA
Step 1: 262 mg of stamping foil PET waste-derived TPA (1.50 mmol, 1.00 equiv.) was added to a suspension of fresh, PBS-washed E. coli_pPCA4_pCAT1 cells (OD600 = 30) in 50 ml of M9-glucose (2%) in a 500 ml Erlenmeyer flask and incubated at 21 °C, 220 rpm for 24 h. Step 2: the resulting suspension was centrifuged at 3,000g, 4 °C for 15 min and the supernatant was transferred to a 50 ml Falcon tube containing sodium pyruvate (60 mM, 2 equiv.), ammonium chloride (1 M, 33.3 equiv.), potassium chloride (10 mM), disodium EDTA (3 mM), sodium sulfite (6.8 mM) and pyridoxal phosphate (0.5 mM) before adjusting the pH to 8 using ammonium hydroxide (28% v/v, aq.). The resulting solution was used to resuspend the E. coli_pPCA4_pCAT1 cell pellet and then a fresh, PBS-washed (1× culture volume) E. coli_pFnTPL cell pellet (post-expression cells OD600 = 30). The resulting suspension was incubated at 21 °C, 220 rpm for 3 h (l-DOPA analytical yield of 84%) before being quenched by acidification with HCl (6 M, aq.) and then centrifuged at 3,000g, 4 °C for 15 min. Acetonitrile was added to the resulting supernatant (final concentration of 5% v/v) before being filtered (0.4 µm) and purified by prep HPLC (Teledyne LABS ACCQPrep HP150, RediSep C18, 5 µm, 18.9 ml min−1, isocratic 5:95 acetonitrile + 0.1% TFA:H2O + 0.1% TFA). Fractions containing l-DOPA were pooled and the organics were removed by rotary evaporator before lyophilization to reveal l-DOPA as the TFA salt (193 mg, 0.62 mmol, 41%).
Plastic waste hydrolysis
Chemical: PET plastic bottle waste and uncoated PET HSF waste was cleaned with 70% ethanol and cut into ~2 cm2 segments. PET fragments (10 g or 20 g) were added to a mixture of 72 ml of ethanol and 288 ml of NaOH solution (10% w/v) and stirred at 90 °C for 10 h at reflux. The reaction was filtered under vacuum before the filtrate was acidified using 100 ml of HCl solution (37% aq.). The resulting white powder was collected, dried by vacuum filtration and used without further purification (Supplementary Table 5 and Supplementary Fig. 17).
Biological: PET food packaging (10 mg, cryo-milled and sorted to a sieve size of 4 mm) was incubated with purified LCCICCM (3 mg of protein per g of PET waste) in 100 mM potassium phosphate buffer (pH 8.0), at 72 °C and 450 rpm for 18 h. The resulting TPA was assessed by HPLC (Supplementary Table 2).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data supporting the findings of this study are available from the article and its Supplementary Information. Source data are provided with this paper.
References
-
Zhao, X., Xiong, L., Zhang, M. & Bai, F. Towards efficient bioethanol production from agricultural and forestry residues: exploration of unique natural microorganisms in combination with advanced strain engineering. Bioresour. Technol. 215, 84–91 (2016).
-
Salvachúa, D., Karp, E. M., Nimlos, C. T., Vardon, D. R. & Beckham, G. T. Towards lignin consolidated bioprocessing: simultaneous lignin depolymerization and product generation by bacteria. Green Chem. 17, 4951–4967 (2015).
-
Kirby, J. et al. Further engineering of R. toruloides for the production of terpenes from lignocellulosic biomass. Biotechnol. Biofuels 14, 101 (2021).
-
Ma, X. et al. Upcycling chitin-containing waste into organonitrogen chemicals via an integrated process. Proc. Natl Acad. Sci. USA 117, 7719–7728 (2020).
-
Suitor, J. T., Varzandeh, S. & Wallace, S. One-pot synthesis of adipic acid from guaiacol in Escherichia coli. ACS Synth. Biol. 9, 2472–2476 (2020).
-
Overhage, J., Steinbüchel, A. & Priefert, H. Harnessing eugenol as a substrate for production of aromatic compounds with recombinant strains of Amycolatopsis sp. HR167. J. Biotechnol. 125, 369–376 (2006).
-
Müller, R. J., Schrader, H., Profe, J., Dresler, K. & Deckwer, W. D. Enzymatic degradation of poly(ethylene terephthalate): rapid hydrolyse using a hydrolase from T. fusca. Macromol. Rapid Commun. 26, 1400–1405 (2005).
-
Yoshida, S. et al. A bacterium that degrades and assimilates poly(ethylene terephthalate). Science 351, 1196–1199 (2016).
-
Bell, E. L. et al. Directed evolution of an efficient and thermostable PET depolymerase. Nat. Catal. 5, 673–681 (2022).
-
Lu, H. et al. Machine learning-aided engineering of hydrolases for PET depolymerization. Nature 604, 662–667 (2022).
-
Sadler, J. C. & Wallace, S. Microbial synthesis of vanillin from waste poly(ethylene terephthalate). Green Chem. 23, 4665–4672 (2021).
-
Valenzuela-Ortega, M., Suitor, J. T., White, M. F. M., Hinchcliffe, T. & Wallace, S. Microbial upcycling of waste PET to adipic acid. ACS Cent. Sci. 9, 2057–2063 (2023).
-
Johnson, N. W. et al. A biocompatible Lossen rearrangement in Escherichia coli. Nat. Chem. 17, 1020–1026 (2025).
-
Kim, H. T. et al. Biological valorization of poly(ethylene terephthalate) monomers for upcycling waste PET. ACS Sustain. Chem. Eng. 7, 19396–19406 (2019).
-
Kim, H. T. et al. Chemo-biological upcycling of poly(ethylene terephthalate) to multifunctional coating materials. ChemSusChem 14, 4251–4259 (2021).
-
Armijo-Galdames, B. O. & Sadler, J. C. One-pot biosynthesis of acetone from waste poly(hydroxybutyrate). ACS Sustain. Chem. Eng. 12, 7748–7756 (2024).
-
Bornscheuer, U. et al. Chemo-enzymatic depolymerization of functionalized low-molecular-weight polyethylene. Angew. Chem. 63, e202415012 (2024).
-
Sullivan, K. P. et al. Mixed plastics waste valorization through tandem chemical oxidation and biological funneling. Science 378, 207–211 (2022).
-
Sadler, J. C., Dennis, J. A., Johnson, N. W. & Wallace, S. Interfacing non-enzymatic catalysis with living microorganisms. RSC Chem. Biol. 2, 1073–1083 (2021).
-
Rabot, C. et al. Polystyrene upcycling into fungal natural products and a biocontrol agent. J. Am. Chem. Soc. 145, 5222–5230 (2023).
-
Koyanagi, T. et al. Effective production of 3,4-dihydroxyphenyl-L-alanine (L-DOPA) with Erwinia herbicola cells carrying a mutant transcriptional regulator TyrR. J. Biotechnol. 115, 303–306 (2005).
-
Savica, R., Grossardt, B. R., Bower, J. H., Eric Ahlskog, J. & Rocca, W. A. Time trends in the incidence of Parkinson disease. JAMA Neurol. 73, 981–989 (2016).
-
Tysnes, O. B. & Storstein, A. Epidemiology of Parkinson’s disease. J. Neural Transm. 124, 901–905 (2017).
-
Soares, A. R. et al. The role of L-DOPA in plants. Plant Signal. Behav. 9, e28275 (2014).
-
Nakagawa, A. et al. A bacterial platform for fermentative production of plant alkaloids. Nat. Commun. 2, 326 (2011).
-
Park, H. S., Lee, J. Y. & Kim, H. S. Production of L-DOPA(3,4-dihydroxyphenyl-L-alanine) from benzene by using a hybrid pathway. Biotechnol. Bioeng. 58, 339–343 (1998).
-
Kumagai, H., Katayama, T., Koyanagi, T. & Suzuki, H. Research overview of L-DOPA production using a bacterial enzyme, tyrosine phenol-lyase. Proc. Japan Acad. B 99, 75–101 (2023).
-
Krishnaveni, R., Rathod, V., Thakur, M. S. & Neelgund, Y. F. Transformation of L-tyrosine to L-dopa by a novel fungus, Acremonium rutilum, under submerged fermentation. Curr. Microbiol. 58, 122–128 (2009).
-
Wei, T., Cheng, B. Y. & Liu, J. Z. Genome engineering Escherichia coli for L-DOPA overproduction from glucose. Sci. Rep. 6, 30080 (2016).
-
Breitel, D. et al. Metabolic engineering of tomato fruit enriched in L-DOPA. Metab. Eng. 65, 185–196 (2021).
-
Patrauchan, M. A. et al. Catabolism of benzoate and phthalate in Rhodococcus sp. strain RHA1: redundancies and convergence. J. Bacteriol. 187, 4050–4063 (2005).
-
Salvador, M. et al. Microbial genes for a circular and sustainable bio-PET economy. Genes 10, 373 (2019).
-
Schweigert, N., Zehnder, A. J. B. & Eggen, R. I. L. Chemical properties of catechols and their molecular modes of toxic action in cells, from microorganisms to mammals. Environ. Microbiol. 3, 81–91 (2001).
-
Li, Q., Aubrey, M. T., Christian, T. & Freed, B. M. Differential inhibition of DNA synthesis in human T cells by the cigarette tar components hydroquinone and catechol. Fundam. Appl. Toxicol. 38, 158–165 (1997).
-
Kobayashi, T. et al. 3,5-Dihydroxybenzoic acid as a potent inhibitor of tyrosine phenol-lyase decreases fecal phenol levels in mice. J. Med. Chem. 68, 8786–8795 (2025).
-
Tang, X. L. et al. Process development for efficient biosynthesis of l-DOPA with recombinant Escherichia coli harboring tyrosine phenol lyase from Fusobacterium nucleatum. Bioprocess. Biosyst. Eng. 41, 1347–1354 (2018).
-
Zhang, X. et al. Endogenous 3,4-dihydroxyphenylalanine and dopaquinone modifications on protein tyrosine: links to mitochondrially derived oxidative stress via hydroxyl radical. Mol. Cell. Prot. 9, 1199–1208 (2010).
-
Valenzuela-Ortega, M. & French, C. Joint universal modular plasmids (JUMP): a flexible vector platform for synthetic biology. Synth. Biol. 6, ysab003 (2021).
Acknowledgements
B.R. acknowledges a PhD studentship from the Industrial Biotechnology Innovation Centre (IBioIC). S.W. acknowledges a Future Leaders Fellowship from UKRI (MR/S033882/1), Sustainable Manufacturing grant from EPSRC (EP/W019000/1), Engineering Biology Mission Hub grant from BBSRC (BB/Y007972/1) and Sustainable Manufacturing Hub grant (UKRI1891). C.W.W. acknowledges a sLOLA grant from BBSRC (BB/X003027/1). C.U. acknowledges NSRF funding via the Research and Innovation Acceleration Agency for Competitiveness and Area Development (RCAD) (Program Management Unit for Technology and Innovation for Future Industries (PMU-B): Brainpower for Future Industries; grant number B38G690002). B.E. acknowledges research assistant and studentship funds from VISTEC. We thank G. Leung and T. Hinchcliffe (Impact Solutions) for insightful discussions, B. French (API Foilmakers) for providing HSF samples, C. P. Lilly and A. Molnar for providing a strain of C. reinhardtii CC1690 and experimental assistance and R. Cox (C-Source Renewables) for supplying bread waste glucose syrups.
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Sustainability thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Royer, B., Era, Y., Valenzuela-Ortega, M. et al. Microbial upcycling of plastic waste to levodopa. Nat Sustain (2026). https://doi.org/10.1038/s41893-026-01785-z
-
Received:
-
Accepted:
-
Published:
-
Version of record:
-
DOI: https://doi.org/10.1038/s41893-026-01785-z